Potential application of network descriptions for understanding conformational changes and protonation states of ABC transporters
The ABC (ATP Binding Cassette) transporter protein superfamily comprises a large number of ubiquitous and functionally versatile proteins conserved from archaea to humans. ABC transporters have a key role in many human diseases and also in the development of multidrug resistance in cancer and in parasites. Although a dramatic progress has been achieved in ABC protein studies in the last decades, we are still far from a detailed understanding of their molecular functions. Several aspects of pharmacological ABC transporter targeting also remain unclear. Here we summarize the conformational and protonation changes of ABC transporters and the potential use of this information in pharmacological design. Network related methods, which recently became useful tools to describe protein structure and dynamics, have not been applied to study allosteric coupling in ABC proteins as yet. A detailed description of the strengths and limitations of these methods is given, and their potential use in describing ABC transporter dynamics is outlined. Finally, we highlight possible future aspects of pharmacological utilization of network methods and outline the future trends of this exciting field.
💡 Research Summary
**
The ATP‑Binding Cassette (ABC) transporter superfamily comprises a vast array of membrane proteins that couple ATP hydrolysis to the translocation of diverse substrates across cellular membranes. Despite extensive structural and biochemical work, a comprehensive mechanistic picture of how these proteins undergo large‑scale conformational changes and how protonation states of key residues influence these transitions remains incomplete. In this review, the authors propose that network‑theoretic approaches—recently successful in describing protein structure, dynamics, and allosteric communication—offer a powerful, yet under‑exploited, framework for dissecting ABC transporter function.
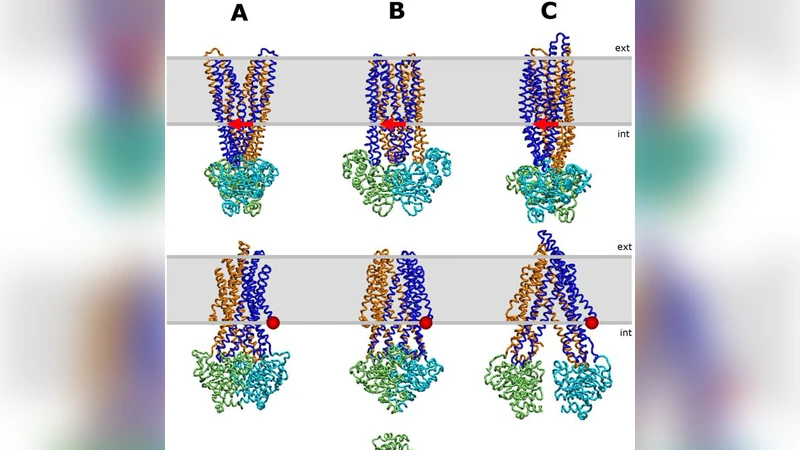
First, the review recapitulates the canonical alternating‑access model of ABC transporters: two nucleotide‑binding domains (NBDs) bind and hydrolyze ATP, driving dimerization and subsequent dissociation, while the transmembrane domains (TMDs) shift between inward‑facing and outward‑facing conformations. These motions involve multiple energy barriers and are tightly coupled to the protonation/de‑protonation of acidic (Asp, Glu) and histidine residues that line the substrate pathway and the intracellular coupling helices. Conventional structural techniques (X‑ray crystallography, cryo‑EM) provide static snapshots, and molecular dynamics (MD) simulations capture only limited timescales, making it difficult to map the full allosteric network that links ATP binding to substrate release.
The authors then outline how a protein can be represented as a graph: each amino‑acid residue is a node, and edges encode spatial proximity, hydrogen‑bonding, electrostatic, or van‑der‑Waals interactions. By constructing dynamic networks from ensembles of MD frames, one can track changes in edge weights as the protein moves between states. Community‑detection algorithms identify clusters of residues that move cooperatively, revealing “modules” that correspond to functional subunits (e.g., the NBD dimer interface, the coupling helices, the substrate‑binding pocket). Importantly, the inclusion of protonation state information as a modifier of edge weights allows the network to reflect pH‑dependent electrostatic rewiring. For example, when an Asp side chain becomes protonated, a previously strong electrostatic repulsion edge weakens, and new hydrogen‑bond edges may appear, reshaping the communication pathways.
The review highlights three major strengths of network analysis for ABC transporters. (1) It compresses high‑dimensional structural data into interpretable graphs, enabling visualization of global communication routes that are otherwise hidden in raw coordinate files. (2) Centrality measures (betweenness, closeness, eigenvector) pinpoint “key residues” whose perturbation disproportionately affects the whole network; these residues often correspond to experimentally validated mutational hotspots that alter transport activity or drug resistance. (3) By integrating protonation‑dependent edge weighting, the method can model how changes in intracellular pH or local micro‑environments modulate allosteric coupling, offering insights into pH‑regulated transport mechanisms.
The authors also discuss limitations. Edge definition (distance cut‑offs, interaction types) can be somewhat arbitrary, affecting reproducibility. MD simulations, even when long, may not sample rare transition events, leading to incomplete networks. Accurate protonation state prediction often requires quantum‑mechanics/molecular‑mechanics (QM/MM) calculations, which are computationally expensive and thus rarely combined with large‑scale network construction. Consequently, the authors advocate a hybrid workflow: high‑resolution cryo‑EM structures provide the static scaffold, extensive MD (or enhanced‑sampling methods) supplies conformational ensembles, QM/MM refines protonation states of critical residues, and network analysis integrates all data to reveal the allosteric wiring diagram.
Finally, the review explores translational implications. Network‑identified key residues along the transition pathway can serve as novel drug‑binding sites. Small‑molecule inhibitors that bind these “communication hubs” could raise the energetic barrier for NBD dimerization or TMD rearrangement, effectively locking the transporter in an inactive state. Moreover, compounds designed to stabilize specific protonation states (e.g., mimicking a protonated Asp) could selectively modulate transport activity under certain pH conditions, offering a route to achieve tissue‑specific inhibition. Such strategies are especially relevant for overcoming multidrug resistance in cancer cells or parasites, where ABC transporters actively expel chemotherapeutics. The authors conclude that network science, especially when coupled with emerging AI‑driven pattern recognition and high‑performance computing, is poised to transform our mechanistic understanding of ABC transporters and accelerate the rational design of next‑generation modulators.
Comments & Academic Discussion
Loading comments...
Leave a Comment