Stability of graph communities across time scales
The complexity of biological, social and engineering networks makes it desirable to find natural partitions into communities that can act as simplified descriptions and provide insight into the structure and function of the overall system. Although community detection methods abound, there is a lack of consensus on how to quantify and rank the quality of partitions. We show here that the quality of a partition can be measured in terms of its stability, defined in terms of the clustered autocovariance of a Markov process taking place on the graph. Because the stability has an intrinsic dependence on time scales of the graph, it allows us to compare and rank partitions at each time and also to establish the time spans over which partitions are optimal. Hence the Markov time acts effectively as an intrinsic resolution parameter that establishes a hierarchy of increasingly coarser clusterings. Within our framework we can then provide a unifying view of several standard partitioning measures: modularity and normalized cut size can be interpreted as one-step time measures, whereas Fiedler’s spectral clustering emerges at long times. We apply our method to characterize the relevance and persistence of partitions over time for constructive and real networks, including hierarchical graphs and social networks. We also obtain reduced descriptions for atomic level protein structures over different time scales.
💡 Research Summary
The paper introduces a novel framework for evaluating the quality of graph partitions by measuring their “stability,” defined through the clustered autocovariance of a Markov process running on the graph. The authors start by formalizing a discrete‑time random walk on a network with transition matrix P and stationary distribution π. For any given partition C, they compute the probability that a random walker, after t steps, remains within the same community, subtract the corresponding probability under a null model of independent assignment, and define the resulting quantity R(t, C) as the stability of the partition at Markov time t. Positive stability indicates that the partition captures more structure than random at that time scale.
A key insight is that the Markov time t acts as an intrinsic resolution parameter. At short times the walk explores only local neighborhoods, so fine‑grained, densely connected subgraphs dominate the stability. As t increases, the walk mixes globally, and the stability reflects coarser, more global structures. Consequently, each time scale yields its own optimal partition, and the evolution of R(t, C) across t provides a natural hierarchy of community resolutions.
The authors demonstrate that several classic quality functions are special cases of this framework. When t = 1, stability reduces exactly to Newman–Girvan modularity, showing that modularity corresponds to a one‑step random‑walk view of the network. In the limit t → ∞, the dominant term of the autocovariance is governed by the second eigenvector of the graph Laplacian (the Fiedler vector), thus recovering spectral clustering based on the normalized cut. Normalized cut itself emerges as a one‑step measure analogous to modularity but with a different null model. In this way, the Markov time unifies modularity, normalized cut, and spectral methods under a single dynamical perspective.
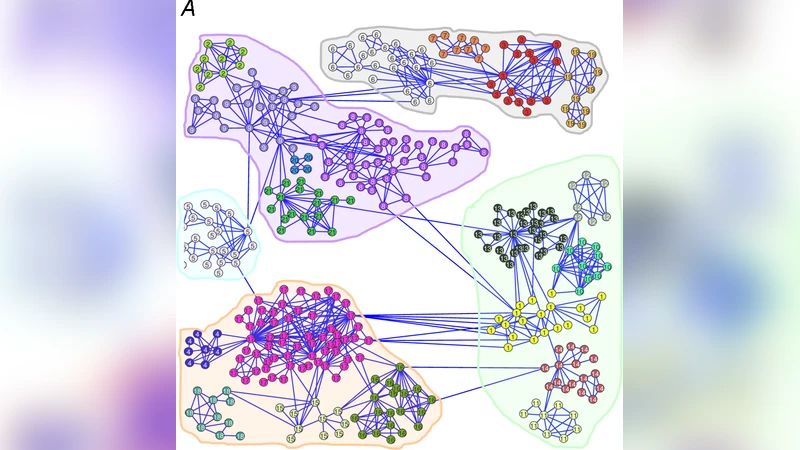
To validate the approach, the paper presents three families of experiments. First, synthetic hierarchical graphs are constructed with multiple nested community levels. By plotting stability versus t for each candidate partition, the authors show that each hierarchical level is most stable over a distinct time interval, effectively identifying the “lifespan” of each community scale. Second, real‑world social networks (e.g., a Facebook friendship graph) are analyzed. Certain social groups (such as university cohorts) exhibit high stability over extended time windows, indicating that these groups are robust structural features of the network dynamics. Third, the method is applied to protein‑level interaction graphs derived from atomic coordinates. Short‑time stability highlights local residue clusters, while long‑time stability reveals domain‑scale modules that correspond closely to known functional regions of the protein, demonstrating the method’s relevance for structural biology.
From a computational standpoint, the authors address scalability by avoiding explicit matrix exponentiation. They employ Krylov subspace techniques (Lanczos or Arnoldi iterations) to approximate P^t·π efficiently for large sparse graphs, and they integrate the stability measure into existing community‑detection heuristics (e.g., the Louvain algorithm) by iteratively optimizing R(t, C) at successive time scales. This yields a practical pipeline that can extract multi‑scale partitions from networks with millions of nodes.
In summary, the paper proposes a dynamic, time‑dependent quality function for graph partitions that simultaneously quantifies partition quality and provides a built‑in resolution parameter. By interpreting community detection as a problem of preserving Markov‑process autocovariance, the framework unifies several widely used static measures, explains their limitations, and extends them to a continuous spectrum of scales. The empirical results on synthetic hierarchies, social graphs, and protein structures illustrate that stability not only ranks partitions at each scale but also reveals the temporal persistence of structural features, offering a powerful tool for multi‑scale network analysis across disciplines.
Comments & Academic Discussion
Loading comments...
Leave a Comment