The Geometry of Morphogenesis and the Morphogenetic Field Concept
The process of morphogenesis, which can be defined as an evolution of the form of an organism, is one of the most intriguing mysteries in the life sciences. It is clear, that gene expression patterns cannot explain the development of the precise geometry of an organism and its parts in space. Here, we suggest a set of postulates and possible approaches for discovering the correspondence between molecular biological information and its realization in a given geometry of an organism in space-time. First, we suggest that the geometry of the organism and its parts is coded by a molecular code located on the cell surfaces in such a way that, with each cell, there can be associated a corresponding matrix, containing this code. As a particular model, we propose coding by several types of oligosaccharide residues of glycoconjugates. Second, we provide a notion of cell event, and suggest a description of development as a tree of cell events, where under cell event we understand the changing of cell state, e.g. the processes of cell division, cell growth/death, shift or differentiation. Next we suggest describing these cell motion laws using the notion of a “morphogenetic field”, meaning by the field an object in an “event space” over a “cell space”, which govern the transformation of the coded biological information into an instructive signal for a cell event for a given cell, depending on the position of a cell in the developing embryo. The matrix on a cell surface will be changed after each cell event according to the rule(s) dictated by the morphogenetic field of an organism. Finally, we provide some ideas on the connection between morphogenetic code on a cell surface, cell motion law(s) and the geometry of an embryo.
💡 Research Summary
The paper tackles one of the most persistent puzzles in developmental biology: how the precise three‑dimensional geometry of an organism emerges from molecular information. The authors argue that gene expression patterns alone cannot account for the spatially intricate forms observed during morphogenesis and propose a new conceptual framework that links molecular codes on cell surfaces to the global geometry of the embryo through a “morphogenetic field.”
First, they postulate the existence of a morphogenetic code embedded in the plasma membrane of every cell. This code is represented mathematically as a matrix that can store high‑dimensional information about the cell’s positional identity and its future developmental potential. As a concrete biochemical candidate, the authors suggest the pattern of oligosaccharide residues on glycoconjugates (glycoproteins, glycolipids, etc.). Because carbohydrate structures can vary combinatorially, are displayed on the extracellular side of the membrane, and are known to mediate cell‑cell recognition, they are well suited to act as a “surface barcode.”
Second, the paper defines a cell event as any discrete change in a cell’s state: division, growth, apoptosis, migration, or differentiation. The entire developmental process is then modeled as a tree of cell events, where each node corresponds to a specific event and edges encode the lineage relationships between parent and daughter cells. This representation captures both the temporal sequence of events and the branching architecture of the organism.
Third, the authors introduce the notion of a morphogenetic field. Unlike classic morphogen gradients, this field is an abstract object defined over an “event space” that overlays the physical “cell space.” For any given cell at a given time, the field receives as input the cell’s surface code matrix together with the spatial context (positions of neighboring cells, external mechanical or chemical cues). It then outputs an instructional signal that determines which cell event should be executed next. In mathematical terms, the field can be described as a vector or tensor field whose components are functions of the local code matrices and of global parameters such as embryo size, curvature, or boundary conditions.
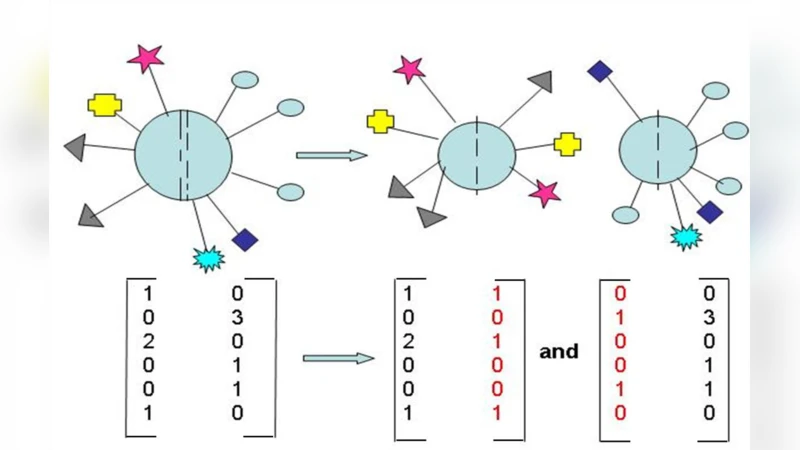
The field also dictates how the surface code is updated after each event. For example, when a cell divides, the parent’s matrix is transformed by a rule (e.g., multiplication by a transformation matrix, addition of a bias vector) to generate the matrices of the two daughter cells. These transformation rules can be expressed as differential equations or discrete update maps, allowing the whole system to be simulated computationally from an initial condition (the fertilized egg’s code) to the final organ geometry.
To bridge theory and experiment, the authors outline three complementary strategies:
- High‑resolution glycomics – using mass spectrometry, NMR, or lectin‑microarray profiling to map the oligosaccharide composition of individual cells in a developing embryo.
- Genetic manipulation of carbohydrate biosynthesis – employing CRISPR/Cas9 to knock out or over‑express specific glycosyltransferases, thereby creating predictable alterations in the surface code and observing the resulting morphological phenotypes.
- Model validation – constructing computational simulations based on the proposed field equations and comparing the predicted tissue shapes, branching patterns, and timing of events with live‑imaging data (e.g., light‑sheet microscopy) of embryos.
The authors argue that this “code‑field‑form” triad supersedes the classic morphogen‑gradient model, which relies on diffusion‑reaction dynamics to generate concentration fields that cells read out. While morphogen gradients explain many patterning phenomena, they struggle to account for the high‑dimensional, non‑linear mapping required to produce complex, asymmetric organs. By embedding a rich, combinatorial code directly on each cell and letting a global field translate that code into context‑dependent actions, the framework can, in principle, generate the full repertoire of embryonic shapes.
Several challenges are acknowledged. Translating the complex carbohydrate patterns into a mathematically tractable matrix representation remains non‑trivial; dimensionality reduction or feature extraction methods will be needed. Defining the morphogenetic field in a way that is both biologically plausible and mathematically solvable requires careful choice of functional forms and parameters. Finally, reconstructing the complete tree of cell events for a whole organism demands large‑scale, high‑temporal‑resolution lineage tracing, which is still at the frontier of imaging technology.
In conclusion, the paper presents a bold, interdisciplinary proposal that re‑conceptualizes morphogenesis as a process governed by surface‑encoded molecular information, a globally acting morphogenetic field, and a deterministic set of cell‑state transitions. If the suggested experimental pipelines can be realized and the mathematical models refined, this approach could provide a unified language for linking genotype, cellular biochemistry, and organismal geometry, opening new avenues for predictive developmental biology and engineered tissue design.
Comments & Academic Discussion
Loading comments...
Leave a Comment