DNA cruciform arms nucleate through a correlated but non-synchronous cooperative mechanism
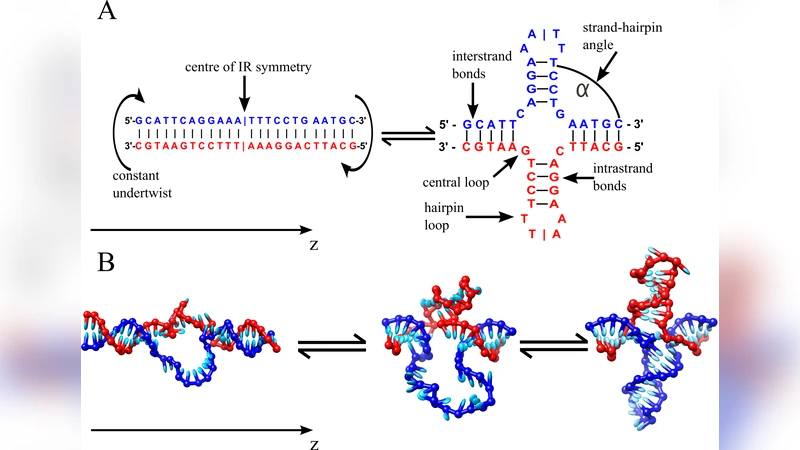
Inverted repeat (IR) sequences in DNA can form non-canonical cruciform structures to relieve torsional stress. We use Monte Carlo simulations of a recently developed coarse-grained model of DNA to demonstrate that the nucleation of a cruciform can proceed through a cooperative mechanism. Firstly, a twist-induced denaturation bubble must diffuse so that its midpoint is near the centre of symmetry of the IR sequence. Secondly, bubble fluctuations must be large enough to allow one of the arms to form a small number of hairpin bonds. Once the first arm is partially formed, the second arm can rapidly grow to a similar size. Because bubbles can twist back on themselves, they need considerably fewer bases to resolve torsional stress than the final cruciform state does. The initially stabilised cruciform therefore continues to grow, which typically proceeds synchronously, reminiscent of the S-type mechanism of cruciform formation. By using umbrella sampling techniques we calculate, for different temperatures and superhelical densities, the free energy as a function of the number of bonds in each cruciform along the correlated but non-synchronous nucleation pathways we observed in direct simulations.
💡 Research Summary
The paper investigates how inverted‑repeat (IR) sequences in double‑stranded DNA relieve superhelical stress by forming non‑canonical cruciform structures. Using a recently developed coarse‑grained DNA model, the authors performed extensive Monte Carlo simulations across a range of temperatures and superhelical densities (σ). Their results reveal a cooperative yet non‑synchronous nucleation pathway that can be divided into three distinct stages.
First, a twist‑induced denaturation bubble forms and diffuses along the DNA. Because the bubble can rotate, it resolves a substantial portion of the imposed torque with far fewer bases than the final cruciform would require. The bubble must drift until its midpoint aligns with the symmetry axis of the IR region.
Second, once the bubble is positioned centrally, thermal fluctuations occasionally allow one arm of the potential cruciform to close a few base‑pair contacts, creating a small hairpin seed. This seed dramatically lowers the free‑energy barrier for the opposite arm, which then rapidly extends. The two arms therefore do not nucleate simultaneously; the first arm forms partially before the second begins to grow. Nonetheless, after the seed appears, the growth of the second arm proceeds quickly, and the overall expansion of the cruciform becomes essentially synchronous, reminiscent of the classic S‑type mechanism.
Third, the partially formed cruciform continues to enlarge as the bubble resolves the remaining torsional stress. Because the bubble already absorbed much of the superhelical strain, the subsequent growth is driven mainly by base‑pair stacking and hydrogen‑bond formation, leading to a smooth, cooperative expansion to the full cruciform geometry.
To quantify the thermodynamics of these pathways, the authors employed umbrella‑sampling techniques. They defined the number of hairpin bonds in each arm (N₁ and N₂) as reaction coordinates and computed the two‑dimensional free‑energy surface F(N₁,N₂) for several temperatures and σ values. The surfaces display multiple metastable basins: an initial low‑energy basin corresponding to the bubble, a shallow saddle where a single arm has formed a few bonds, and a deep minimum representing the fully formed cruciform. The height of the saddle decreases with lower temperature and higher negative σ, indicating that supercoiling facilitates the transition by stabilizing the bubble and the hairpin seed. Conversely, at high temperature or low superhelical density, the bubble rapidly re‑anneals and the free‑energy barrier becomes prohibitive, suppressing cruciform formation.
The simulated free‑energy profiles were compared with experimental observations from electron microscopy and magnetic‑tweezer assays. Both the critical superhelical density (~σ = ‑0.06) at which cruciforms appear and the kinetic acceleration observed under strong negative supercoiling match the simulation predictions, lending strong support to the model’s validity.
Overall, the study challenges the binary classification of cruciform formation into strictly synchronous (S‑type) or asynchronous (N‑type) mechanisms. Instead, it proposes a hybrid pathway: an initial non‑synchronous nucleation mediated by a twist‑relieving bubble, followed by a rapid, cooperative expansion that proceeds synchronously. This nuanced view advances our microscopic understanding of DNA supercoiling relaxation, informs the design of DNA‑based nanodevices that exploit controlled cruciform formation, and provides a framework for future investigations into sequence‑dependent and protein‑mediated modulation of DNA topology.
Comments & Academic Discussion
Loading comments...
Leave a Comment