A phylogeny of birds based on over 1,500 loci collected by target enrichment and high-throughput sequencing
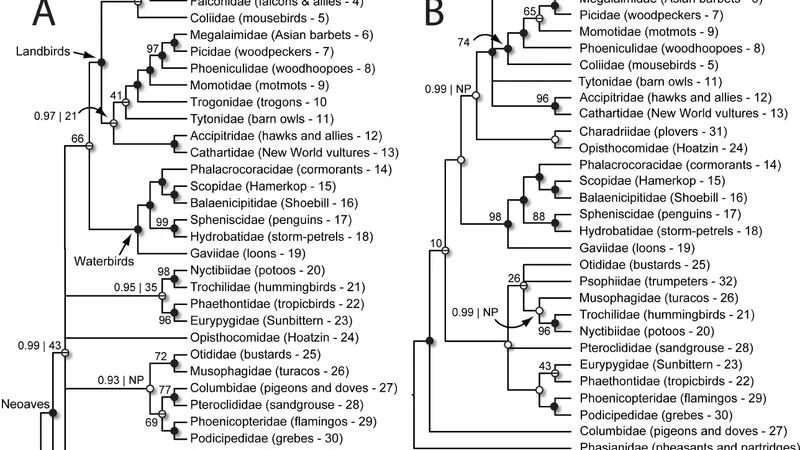
Evolutionary relationships among birds in Neoaves, the clade comprising the vast majority of avian diversity, have vexed systematists due to the ancient, rapid radiation of numerous lineages. We applied a new phylogenomic approach to resolve relationships in Neoaves using target enrichment (sequence capture) and high-throughput sequencing of ultraconserved elements (UCEs) in avian genomes. We collected sequence data from UCE loci for 32 members of Neoaves and one outgroup (chicken) and analyzed data sets that differed in their amount of missing data. An alignment of 1,541 loci that allowed missing data was 87% complete and resulted in a highly resolved phylogeny with broad agreement between the Bayesian and maximum-likelihood (ML) trees. Although results from the 100% complete matrix of 416 UCE loci were similar, the Bayesian and ML trees differed to a greater extent in this analysis, suggesting that increasing from 416 to 1,541 loci led to increased stability and resolution of the tree. Novel results of our study include surprisingly close relationships between phenotypically divergent bird families, such as tropicbirds (Phaethontidae) and the sunbittern (Eurypygidae) as well as between bustards (Otididae) and turacos (Musophagidae). This phylogeny bolsters support for monophyletic waterbird and landbird clades and also strongly supports controversial results from previous studies, including the sister relationship between passerines and parrots and the non-monophyly of raptorial birds in the hawk and falcon families. Although significant challenges remain to fully resolving some of the deep relationships in Neoaves, especially among lineages outside the waterbirds and landbirds, this study suggests that increased data will yield an increasingly resolved avian phylogeny.
💡 Research Summary
The authors address one of the most persistent challenges in avian systematics: resolving the deep‐time relationships among Neoaves, the clade that contains the overwhelming majority of bird diversity. Because Neoaves underwent an ancient, rapid radiation, traditional markers (mitochondrial genes, a handful of nuclear loci) have produced conflicting and poorly supported trees. To overcome these limitations, the study employs a phylogenomic strategy based on ultraconserved elements (UCEs). Using a set of previously designed avian UCE probes, DNA from 32 Neoaves species and a chicken outgroup was enriched, sequenced on an Illumina platform, and assembled. After quality filtering and alignment, two data matrices were constructed: (1) a “relaxed” matrix containing 1,541 loci that is 87 % complete (allowing missing data) and (2) a “strict” matrix of 416 loci that is 100 % complete across all taxa.
Phylogenetic inference was performed with both Bayesian (MrBayes) and maximum‑likelihood (RAxML) methods. The larger matrix yielded highly congruent Bayesian and ML trees, with most deep nodes receiving posterior probabilities and bootstrap values above 95 %. In contrast, the smaller matrix showed greater topological discordance between the two methods and lower support for several deep splits, demonstrating that increasing the number of loci markedly improves tree stability and resolution.
Key findings reaffirm several controversial relationships that have emerged from recent genome‑scale studies. The waterbird clade (Anseriformes + Galliformes) and the landbird clade (the remainder of Neoaves) are both strongly supported as monophyletic. The sister‑group relationship between Passeriformes (songbirds) and Psittaciformes (parrots) receives high statistical support, as does the non‑monophyly of traditional raptorial birds: falcons (Falconidae) and hawks/eagles (Accipitridae) form separate lineages.
The analysis also uncovers unexpected sister‑group relationships that challenge morphology‑based classifications. Tropicbirds (Phaethontidae) are placed as the closest relatives of the sunbittern (Eurypygidae), and bustards (Otididae) are recovered as sister to turacos (Musophagidae). These results suggest that convergent ecological and phenotypic evolution has obscured true genetic affinities.
The authors discuss the implications of missing data, showing that a matrix with 87 % completeness can still provide robust phylogenetic signal, whereas insisting on 100 % completeness may discard valuable information. They acknowledge that several deep nodes—particularly those outside the well‑supported waterbird and landbird clusters—remain ambiguous, likely because the current UCE probe set captures insufficient variation for those lineages.
In conclusion, the study demonstrates that target enrichment of thousands of UCE loci, combined with high‑throughput sequencing, is a powerful approach for resolving the rapid, ancient radiation of Neoaves. By substantially increasing the amount of independent genetic data, the authors achieve a more stable and well‑supported avian phylogeny, corroborate several contentious relationships, and reveal novel sister‑group connections. The work highlights the need for even larger genomic datasets and broader taxon sampling to finally resolve the remaining uncertainties in the avian tree of life.
Comments & Academic Discussion
Loading comments...
Leave a Comment