Improved estimation of anomalous diffusion exponents in single particle tracking experiments
The Mean Square Displacement is a central tool in the analysis of Single Particle Tracking experiments, shedding light on various biophysical phenomena. Frequently, parameters are extracted by performing time-averages on single particle trajectories followed by ensemble averaging. This procedure however, suffers from two systematic errors when applied to particles that perform anomalous diffusion. The first is significant at short time lags and is induced by measurement errors. The second arises from the natural heterogeneity in biophysical systems. We show how to estimate and correct these two errors and improve the estimation of the anomalous parameters for the whole particle distribution. As a consequence we manage to characterize ensembles of heterogeneous particles even for rather short and noisy measurements where regular time averaged mean square displacement analysis fails. We apply this method to both simulations and in vivo measurements of telomere diffusion in 3T3 mouse embryonic fibroblast cells. The motion of telomeres is found to be subdiffusive with an average exponent constant in time. Individual telomere exponents are normally distributed around the average exponent. The proposed methodology has the potential to improve experimental accuracy while maintaining lower experimental costs and complexity.
💡 Research Summary
The paper addresses a fundamental problem in the analysis of single‑particle tracking (SPT) data: the reliable extraction of anomalous diffusion parameters—most notably the anomalous exponent α—from noisy, short trajectories. The authors identify two systematic sources of bias that plague the standard time‑averaged mean‑square‑displacement (tMSD) approach when particles exhibit sub‑diffusive behavior (α < 1).
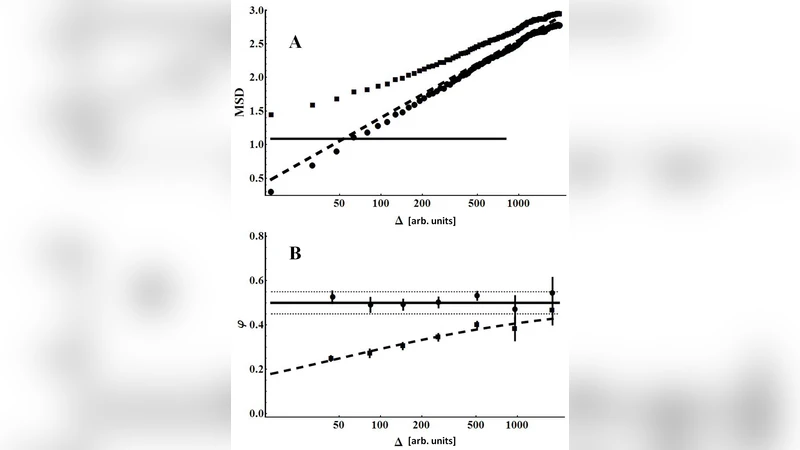
The first bias originates from localization error (σ_loc), an inevitable measurement noise that adds a constant offset (2σ_loc²) to the tMSD at short lag times (Δt). Because the log‑log plot of MSD versus Δt is heavily weighted toward these short lags, the offset artificially depresses the slope and leads to an under‑estimation of α. The authors propose a straightforward correction: independently measure σ_loc (e.g., by tracking immobilized beads) and subtract 2σ_loc² from the raw tMSD before any fitting. This step restores the true physical MSD at all Δt, especially in the regime where the anomalous scaling law holds.
The second bias stems from biological heterogeneity. In realistic cellular environments, each tracked particle experiences a distinct micro‑environment, resulting in a distribution of α and the generalized diffusion coefficient Dα across the population. Conventional practice treats each trajectory independently, fits a line to its log‑log MSD, and then averages the resulting α values. When trajectories are short and noisy, individual fits become highly variable, and the simple average masks the underlying distribution, often yielding misleading ensemble parameters.
To overcome this, the authors introduce a Bayesian mixture‑model framework. They assume that each particle i has its own (α_i, Dα_i) but that these parameters are drawn from a common prior distribution (e.g., a normal distribution for α). Using an Expectation‑Maximization (EM) algorithm, they jointly estimate the hyper‑parameters of the prior (mean μ_α, variance σ_α²) and the posterior expectations of each α_i given the observed trajectory. This hierarchical approach leverages information from the entire dataset, dramatically reducing the variance of individual estimates without requiring longer recordings.
The methodology is validated through extensive simulations. Two scenarios are examined: (1) a homogeneous ensemble with a single α and Dα, and (2) a heterogeneous ensemble where α and Dα follow prescribed normal distributions. In both cases, synthetic localization noise (σ_loc ≈ 20 nm) and varying trajectory lengths (10–100 frames) are added. Compared with the naïve tMSD method, the corrected‑and‑mixed approach reduces the mean absolute error in α by 30–50 % for short trajectories (≤30 frames) and accurately recovers the underlying distribution parameters (μ_α, σ_α).
The authors then apply the technique to real biological data: telomere motion in 3T3 mouse embryonic fibroblasts. Telomeres are fluorescently labeled with TRF1‑GFP and imaged at 20 Hz for 30 s, yielding ~200 trajectories of average length ~30 frames. Raw tMSD analysis suggests an average α≈0.62, heavily biased by short‑lag noise. After measuring σ_loc≈25 nm and applying the offset correction, the log‑log MSD becomes linear over the accessible Δt range. The mixture model then estimates a population‑wide exponent μ_α = 0.78 ± 0.03, with individual α_i distributed normally (σ_α ≈ 0.07). Importantly, the fitted α values show no systematic drift over time, indicating that telomere dynamics are stationary sub‑diffusive rather than time‑varying. The spread in α reflects genuine heterogeneity in the nuclear micro‑environment (e.g., proximity to lamina, chromatin density variations).
In summary, the paper delivers a practical, low‑cost solution to two pervasive pitfalls in SPT analysis: (i) measurement‑induced short‑lag bias and (ii) population heterogeneity. By correcting the tMSD for localization error and embedding the analysis in a Bayesian hierarchical model, researchers can obtain accurate anomalous exponents even from short, noisy recordings. The approach is compatible with existing SPT pipelines and does not demand additional hardware. Potential extensions include monitoring drug‑induced changes in diffusion, dissecting active versus passive transport contributions, and applying the framework to other anomalous processes such as viscoelastic rheology in living cells.
Comments & Academic Discussion
Loading comments...
Leave a Comment