Parallelization in Scientific Workflow Management Systems
Over the last two decades, scientific workflow management systems (SWfMS) have emerged as a means to facilitate the design, execution, and monitoring of reusable scientific data processing pipelines. At the same time, the amounts of data generated in various areas of science outpaced enhancements in computational power and storage capabilities. This is especially true for the life sciences, where new technologies increased the sequencing throughput from kilobytes to terabytes per day. This trend requires current SWfMS to adapt: Native support for parallel workflow execution must be provided to increase performance; dynamically scalable “pay-per-use” compute infrastructures have to be integrated to diminish hardware costs; adaptive scheduling of workflows in distributed compute environments is required to optimize resource utilization. In this survey we give an overview of parallelization techniques for SWfMS, both in theory and in their realization in concrete systems. We find that current systems leave considerable room for improvement and we propose key advancements to the landscape of SWfMS.
💡 Research Summary
The paper provides a comprehensive survey of parallelization techniques in scientific workflow management systems (SWfMS), motivated by the explosive growth of data in fields such as genomics, where daily sequencing output now reaches terabytes. The authors argue that modern SWfMS must evolve to offer native parallel execution, integrate dynamically scalable “pay‑per‑use” compute infrastructures, and adopt adaptive scheduling to make efficient use of distributed resources.
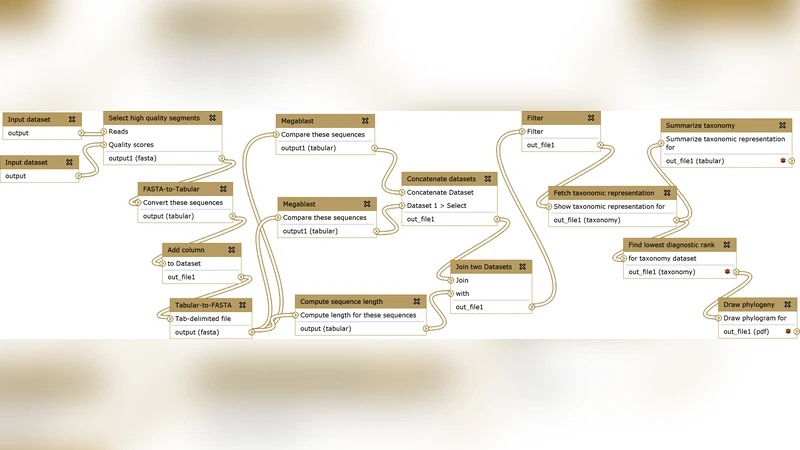
The manuscript begins with a concise overview of scientific workflows: a workflow is modeled as a directed acyclic graph (DAG) whose nodes are tasks that consume input files and produce output files. The authors distinguish three abstraction levels—abstract, concrete, and physical—and explain how workflow planning maps abstract steps to concrete algorithms, while workflow enactment assigns tasks to physical compute resources.
A taxonomy of SWfMS is introduced, separating systems into (1) textual workflow languages (e.g., Pegasus, Swift) that target expert users comfortable with scripts and configuration files, (2) graphical workflow systems (e.g., Taverna, Kepler) that provide drag‑and‑drop interfaces and extensive task libraries, and (3) domain‑specific web portals (e.g., Galaxy, Mobile) that run entirely in a browser and focus on a particular scientific community. This classification frames the subsequent analysis of parallelization support.
The core of the survey is a three‑dimensional taxonomy of parallel execution: (a) types of parallelism, (b) computational infrastructures, and (c) scheduling policies. The three parallelism types are:
-
Task parallelism – independent tasks (nodes) of the DAG are dispatched to different compute nodes. The maximum degree of parallelism can be derived from a static analysis of the DAG, but real‑world execution suffers from heterogeneous task runtimes and data‑dependency bottlenecks, making scheduling non‑trivial.
-
Data parallelism – the same task is applied concurrently to disjoint partitions of its input data. This is especially effective for embarrassingly parallel steps common in next‑generation sequencing (e.g., read alignment, variant calling).
-
Pipeline parallelism – the workflow is treated as a production line; each stage processes a different data batch while upstream and downstream stages run simultaneously, akin to streaming. This approach maximizes throughput for long pipelines with balanced stages.
The infrastructure dimension contrasts traditional cluster/grid environments with modern cloud and container‑orchestrated platforms. Cluster/Grid solutions (e.g., DAGMan, early Pegasus) provide multi‑node and multi‑core execution but often require complex configuration and lack elasticity. Cloud platforms (AWS, Azure, Google Cloud) enable “pay‑per‑use” scaling, object storage (S3), and on‑demand provisioning, yet introduce new concerns such as data transfer costs, latency, and security. The authors also discuss emerging container‑based orchestration (Kubernetes, Docker Swarm) as a way to achieve portable, reproducible deployments across heterogeneous clouds.
Scheduling policies are examined along two axes: static vs. dynamic and centralized vs. decentralized. Static schedulers compute a fixed mapping before execution based on estimated runtimes and resource capacities; they are simple but brittle when actual workloads deviate from predictions. Dynamic schedulers monitor task progress at runtime, reassign tasks, and adapt resource allocation on the fly, thereby handling variability in task duration and resource availability. Most surveyed SWfMS still rely heavily on static approaches, limiting their effectiveness in volatile cloud environments.
The paper then reviews concrete implementations. Taverna and Kepler (graphical systems) excel in usability but provide limited multi‑core support and often require external plugins for distributed execution. Pegasus and Swift (textual systems) offer sophisticated workflow description languages and strong support for grid resources, yet their steep learning curve hampers adoption by domain scientists. KNIME blends a graphical interface with extensible plugins, achieving moderate parallelism. Domain‑specific portals such as Galaxy deliver ready‑made bioinformatics tools and seamless web access, but their architectures are typically tied to a single backend, restricting scalability and cross‑domain reuse.
Throughout the analysis, the authors highlight a persistent trade‑off: systems that prioritize ease of use often sacrifice deep parallel capabilities, while those that expose powerful parallel constructs tend to be less accessible to non‑programmers.
In the final section, the authors outline research directions needed to close this gap:
- Hybrid parallelism models that combine task, data, and pipeline parallelism within a unified execution engine.
- Cloud‑native architectures that leverage containerization, serverless functions, and auto‑scaling groups to provide elastic resources without manual provisioning.
- Machine‑learning‑driven schedulers that predict task runtimes and resource contention, enabling proactive load balancing and cost‑aware scheduling.
- Standardized metadata schemas for workflow description, resource requirements, and provenance, facilitating interoperability across heterogeneous platforms.
- Security‑aware data movement strategies that minimize data exposure while exploiting distributed storage (e.g., encrypted object stores, edge computing).
The authors conclude that addressing these challenges will transform scientific workflows into the de‑facto execution model for large‑scale data‑intensive research, delivering reproducibility, scalability, and cost‑efficiency across disciplines.
Comments & Academic Discussion
Loading comments...
Leave a Comment