A Labeled Graph Kernel for Relationship Extraction
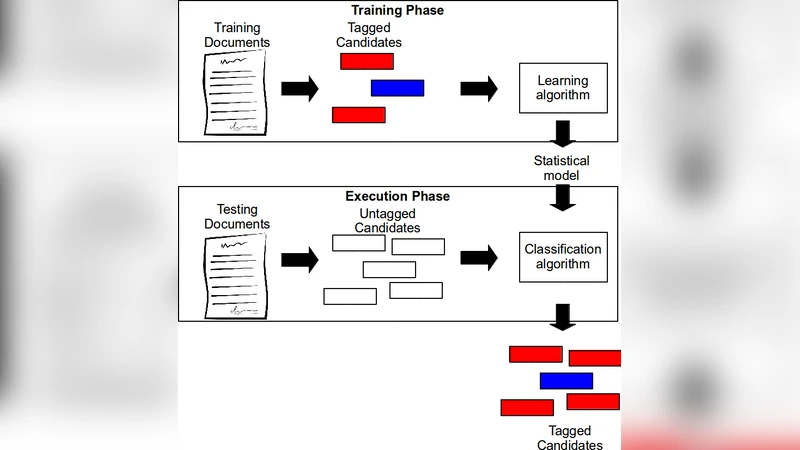
In this paper, we propose an approach for Relationship Extraction (RE) based on labeled graph kernels. The kernel we propose is a particularization of a random walk kernel that exploits two properties previously studied in the RE literature: (i) the words between the candidate entities or connecting them in a syntactic representation are particularly likely to carry information regarding the relationship; and (ii) combining information from distinct sources in a kernel may help the RE system make better decisions. We performed experiments on a dataset of protein-protein interactions and the results show that our approach obtains effectiveness values that are comparable with the state-of-the art kernel methods. Moreover, our approach is able to outperform the state-of-the-art kernels when combined with other kernel methods.
💡 Research Summary
The paper introduces a supervised relationship extraction (RE) method that leverages labeled dependency graphs together with a marginalized random‑walk kernel. The authors start by observing two well‑known hypotheses in the RE literature: (i) the words that lie between two candidate entities—or that connect them in a syntactic representation—are highly informative for determining the existence and type of a relationship, and (ii) combining heterogeneous sources of information within a single kernel can improve classification decisions. To operationalize these ideas, each sentence is transformed into a labeled graph where vertices correspond to tokens enriched with linguistic attributes (POS tag, lemma, capitalization pattern, etc.) and edges encode dependency relations. Two Boolean predicates are defined: isEntity(v), which marks vertices that belong to the candidate entities, and inSP(x), which marks vertices or edges that belong to the shortest path between the two entities. These predicates are incorporated into the kernel as special labels, allowing the kernel to give higher weight to the entities themselves and to the shortest‑path subgraph while still considering the full graph structure.
The underlying similarity measure is the random‑walk kernel for labeled graphs as described by Kashima et al. (2003). In this formulation, similarity between two graphs is computed as the sum over all possible label‑matching random walks, each weighted by transition probabilities and a label‑matching function. By marginalizing over all walks, the kernel implicitly maps graphs into an infinite‑dimensional feature space without explicit enumeration, and the computation reduces to solving a system of linear equations, which is computationally tractable.
The authors evaluate the approach on the AImed corpus, a benchmark dataset of biomedical abstracts annotated with protein‑protein interaction (PPI) relationships. Experiments compare the proposed kernel (both alone and in linear combination with other kernels) against several state‑of‑the‑art graph‑based kernels, including the shortest‑path kernel of Bunescu and Mooney (2005), the shallow‑kernel of Giuliano et al. (2006), and the dual‑graph kernel of Airola et al. (2008). Results show that the standalone labeled‑graph random‑walk kernel achieves performance (precision, recall, F1) comparable to the best existing methods. More importantly, when combined with complementary kernels, the system consistently outperforms each individual kernel, confirming the benefit of multi‑kernel integration.
Key contributions of the work are: (1) a concrete graph representation that enriches dependency structures with entity‑specific and shortest‑path‑specific labels, (2) an adaptation of the marginalized random‑walk kernel to exploit these labels, and (3) empirical evidence that the resulting kernel is both competitive on its own and synergistic when fused with other kernels. The paper also discusses computational considerations: the random‑walk kernel avoids exhaustive path enumeration, leading to lower memory consumption and faster training compared to kernels that explicitly enumerate all paths. However, the overall system remains sensitive to the quality of upstream preprocessing (named‑entity recognition and dependency parsing), as errors in these stages propagate to the graph labels and can degrade kernel similarity scores.
In conclusion, the study demonstrates that a carefully designed labeled‑graph kernel, grounded in well‑established linguistic hypotheses, can serve as a powerful component for relationship extraction, especially in domains where syntactic structure carries rich semantic cues such as biomedical text. Future directions suggested include extending the approach to other relation types, incorporating additional semantic resources (e.g., WordNet synsets, hypernyms), improving robustness to parsing errors, and exploring automatic kernel weight learning for optimal multi‑kernel combinations.
Comments & Academic Discussion
Loading comments...
Leave a Comment