Emulating a gravity model to infer the spatiotemporal dynamics of an infectious disease
Probabilistic models for infectious disease dynamics are useful for understanding the mechanism underlying the spread of infection. When the likelihood function for these models is expensive to evaluate, traditional likelihood-based inference may be computationally intractable. Furthermore, traditional inference may lead to poor parameter estimates and the fitted model may not capture important biological characteristics of the observed data. We propose a novel approach for resolving these issues that is inspired by recent work in emulation and calibration for complex computer models. Our motivating example is the gravity time series susceptible-infected-recovered (TSIR) model. Our approach focuses on the characteristics of the process that are of scientific interest. We find a Gaussian process approximation to the gravity model using key summary statistics obtained from model simulations. We demonstrate via simulated examples that the new approach is computationally expedient, provides accurate parameter inference, and results in a good model fit. We apply our method to analyze measles outbreaks in England and Wales in two periods, the pre-vaccination period from 1944-1965 and the vaccination period from 1966-1994. Based on our results, we are able to obtain important scientific insights about the transmission of measles. In general, our method is applicable to problems where traditional likelihood-based inference is computationally intractable or produces a poor model fit. It is also an alternative to approximate Bayesian computation (ABC) when simulations from the model are expensive.
💡 Research Summary
The paper addresses the formidable computational challenges of fitting spatially explicit infectious‑disease models, focusing on the gravity‑augmented time‑series SIR (TSIR) framework used for measles dynamics in England and Wales. Traditional likelihood‑based inference is prohibitive because each evaluation requires a costly stochastic simulation of the full spatiotemporal process, and naïve point‑estimation methods fail to capture biologically important features such as the distribution of peak incidences and the proportion of disease‑free bi‑weeks.
The authors first attempt a grid‑based Bayesian MCMC that discretizes the parameter space and evaluates the exact likelihood. Simulated experiments reveal that this approach yields biased parameter estimates and model trajectories that do not reproduce the key summary statistics, highlighting the inadequacy of a pure likelihood approach for this model.
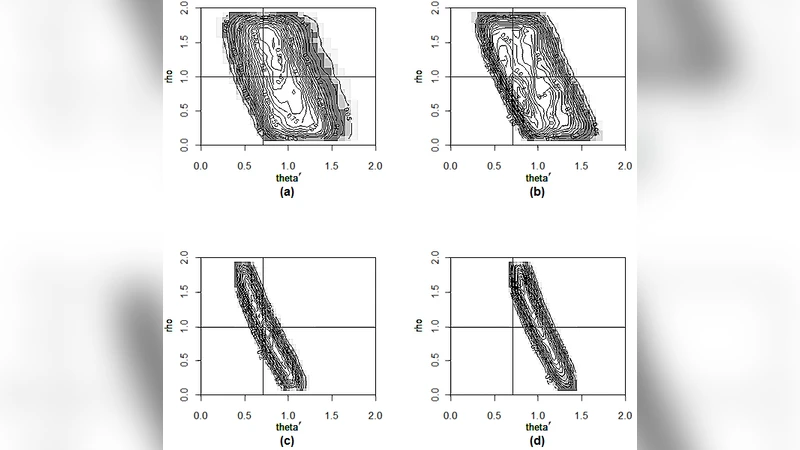
To overcome these limitations, the paper proposes an emulator‑based inference strategy. The core idea is to treat the computational model as a black‑box and build a surrogate using a Gaussian process (GP). A set of carefully chosen summary statistics—city‑wise maximum incidence (M) and the proportion of bi‑weeks with zero cases (P)—are extracted from many model runs across a design of parameter values (θ₀, τ₁, τ₂, ρ). These statistics are selected by domain experts because they encapsulate the epidemiologically relevant dynamics of measles. The GP learns the mapping from the four gravity parameters to the joint distribution of (M, P), providing a fast, probabilistic prediction of the summaries for any new parameter setting.
The GP surrogate is then embedded in a likelihood‑like formulation: the observed summaries are assumed to arise from the GP predictive distribution, and a Metropolis‑Hastings algorithm samples from the resulting posterior. This approach avoids repeated expensive forward simulations while still delivering full Bayesian uncertainty quantification.
Simulation studies demonstrate that the emulator‑based method accurately recovers the true parameters, reproduces the target summary statistics, and yields a substantially better fit than the grid‑based MCMC. The authors also compare their method to Approximate Bayesian Computation (ABC), noting that ABC would be infeasible here because each simulation is already costly, whereas the GP emulator reduces the computational burden dramatically.
Applying the methodology to real measles data, the authors fix the local transmission parameters (βₜ, α) at values estimated in previous work and infer only the gravity parameters. Analyses of the pre‑vaccination era (1944‑1965) and the vaccination era (1966‑1994) reveal several scientific insights:
- Seasonality and holidays – Transmission rates drop markedly during school holidays, confirming the long‑known role of schools as transmission hubs.
- Stability of spatial coupling – The estimated gravity parameters show little change between the two eras, suggesting that the underlying network of infection importation/exportation remained relatively stable despite widespread vaccination.
- Network characterization – Using the posterior draws of the gravity parameters, the authors construct directed infection flow networks among cities, compute degree distributions, and identify key hub cities that consistently act as sources or sinks of measles spread.
These results illustrate that the emulator‑based approach not only yields reliable parameter estimates but also enables biologically meaningful interpretation of complex spatiotemporal disease dynamics.
The paper concludes by emphasizing the broader applicability of the method to any stochastic model with expensive likelihoods and by acknowledging limitations such as the reliance on expert‑chosen summary statistics and the scalability of GP surrogates to very high‑dimensional parameter spaces. Future work is suggested on automated summary selection, sparse GP techniques, and extensions to other infectious diseases.
Comments & Academic Discussion
Loading comments...
Leave a Comment