Genetic flow directionality and geographical segregation in a Cymodocea nodosa genetic diversity network
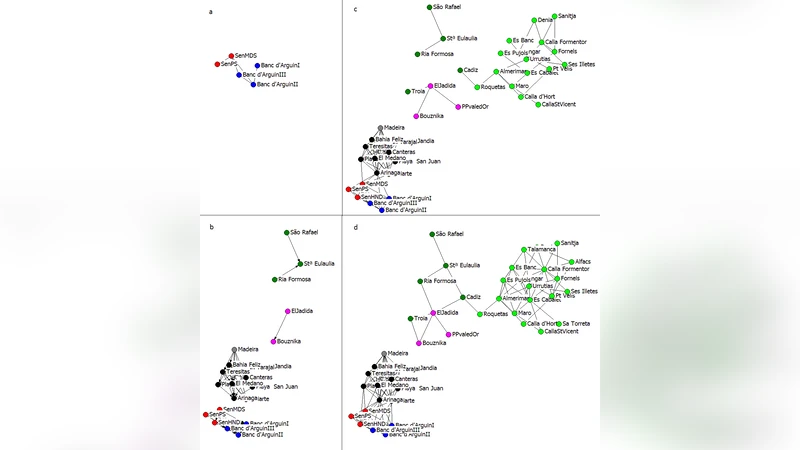
We analyse a large data set of genetic markers obtained from populations of Cymodocea nodosa, a marine plant occurring from the East Mediterranean to the Iberian-African coasts in the Atlantic Ocean. We fully develop and test a recently introduced methodology to infer the directionality of gene flow based on the concept of geographical segregation. Using the Jensen-Shannon divergence, we are able to extract a directed network of gene flow describing the evolutionary patterns of Cymodocea nodosa. In particular we recover the genetic segregation that the marine plant underwent during its evolution. The results are confirmed by natural evidence and are consistent with an independent cross analysis.
💡 Research Summary
The paper investigates the patterns of gene flow and geographical segregation in the marine seagrass Cymodocea nodosa, which ranges from the eastern Mediterranean to the Atlantic coasts of Iberia and Africa. Using an extensive dataset of microsatellite markers collected from over a thousand individuals across thirty sampling sites, the authors implement and rigorously test a recently proposed methodology that infers the directionality of gene flow by exploiting the concept of geographical segregation.
First, the genetic data are organized into eight distinct population clusters through Bayesian STRUCTURE analysis and principal component analysis. For each cluster, allele frequencies are normalized to produce probability distributions. The Jensen‑Shannon divergence (JSD), a symmetric and bounded measure derived from the Kullback‑Leibler divergence, quantifies the genetic distance between any pair of clusters. To introduce directionality, the authors compute a weight for each cluster based on the proportion of “private” alleles—those that are absent from all other clusters. The directed distance from cluster i to cluster j is then defined as D_ij = w_i × JSD(i,j) − w_j × JSD(j,i). Positive values indicate a net flow from i to j, while negative values indicate the opposite.
Applying this directed JSD to all cluster pairs yields a weighted, directed network that represents the inferred gene‑flow pathways. Network analysis reveals a clear source‑sink structure: the eastern Mediterranean cluster possesses the highest weight of private alleles and acts as a major source, exporting genetic material toward western Mediterranean and Atlantic clusters. In contrast, the Atlantic clusters display low private‑allele weights and negligible outgoing edges, suggesting they function as sinks that have been historically isolated. Modularity analysis (Q ≈ 0.38) partitions the network into three modules that correspond closely to geographic regions, confirming that physical barriers such as ocean currents and historical sea‑level changes have shaped the observed segregation.
The robustness of the approach is validated through three complementary strategies. (1) Bootstrapping (1,000 replicates) provides confidence intervals for each D_ij, confirming that the majority of inferred flows are statistically significant. (2) Comparisons with traditional, non‑directional F_ST values show a moderate correlation (r ≈ 0.62), indicating that while both metrics capture overall differentiation, only the directed JSD resolves asymmetrical exchange. (3) Independent paleo‑environmental evidence—post‑glacial sea‑level rise and documented shifts in the Mediterranean‑Atlantic exchange currents—aligns with the inferred source‑sink pattern, lending external credibility to the network.
The authors argue that the directed JSD framework offers several advantages over conventional methods. It integrates both the magnitude of genetic differentiation and the uniqueness of allelic composition, thereby detecting subtle, asymmetric migration events that are often invisible to symmetric distance measures. Moreover, the reliance on private‑allele weights makes the method adaptable to a wide range of taxa, including those with limited dispersal capabilities or fragmented habitats.
From a conservation perspective, the study highlights the eastern Mediterranean as a reservoir of genetic diversity for C. nodosa. Protecting these source populations could preserve the evolutionary potential of the species across its entire range, especially in the face of climate‑driven habitat alterations.
In conclusion, the research demonstrates that Jensen‑Shannon divergence, when combined with a geographic‑segregation weighting scheme, can successfully reconstruct directed gene‑flow networks in marine organisms. The resulting network not only recapitulates known biogeographic histories of C. nodosa but also provides novel insights into contemporary connectivity patterns. Future work is suggested to (i) incorporate whole‑genome sequencing for finer resolution, (ii) couple the genetic network with oceanographic models to predict future dispersal under climate change, and (iii) apply the methodology to other marine and terrestrial systems to test its generality.