Towards a Nano Geometry? Geometry and Dynamics on Nano Scale
This paper applies I.M. Gelfand’s distinction between adequate and non-adequate use of mathematical language in different contexts to the newly opened window of model-based measurements of intracellular dynamics. The specifics of geometry and dynamics on the mesoscale of cell physiology are elaborated - in contrast to the familiar Newtonian mechanics and the more recent, but by now also rather well established quantum field theories. Examples are given originating from the systems biology of insulin secreting pancreatic beta-cells and the mathematical challenges of an envisioned non-invasive control of magnetic nanoparticles.
💡 Research Summary
The paper revisits I.M. Gelfand’s philosophical distinction between “adequate” and “non‑adequate” uses of mathematical language and applies it to the emerging field of model‑based intracellular measurement, focusing on the mesoscopic scale that lies between the realms traditionally described by Newtonian mechanics and quantum field theory. The authors argue that the conventional frameworks—continuum mechanics for macroscopic objects and quantum field theory for sub‑atomic particles—both break down when one attempts to describe phenomena occurring at length scales of tens of nanometers to a few hundred micrometers, the scale at which many essential cellular processes unfold.
The manuscript first defines this “mesoscale” as a regime where the assumptions of continuous media are no longer fully valid, yet quantum coherence effects are negligible. In this regime, cellular membranes exhibit complex curvature, organelles are topologically intertwined, and the cytoskeleton undergoes rapid remodeling. These geometric features cannot be captured adequately by simple differential equations; instead, topological invariants such as Betti numbers, homology groups, and curvature‑dependent tensors become the natural language.
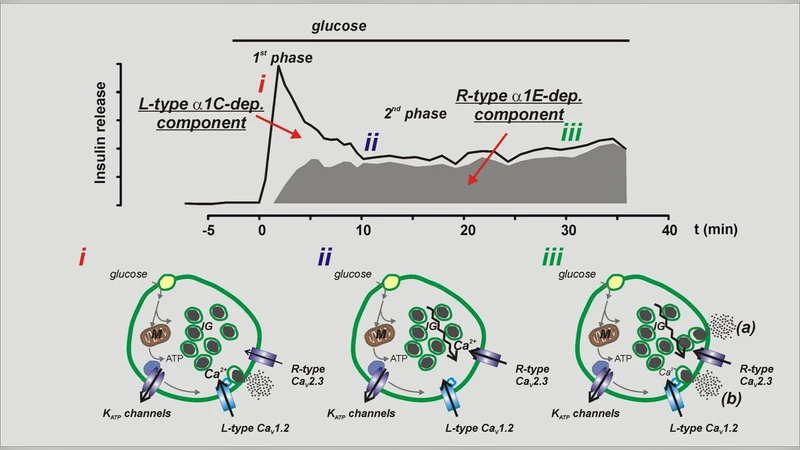
Using pancreatic β‑cells as a concrete example, the authors dissect the insulin secretion pathway. Traditional models treat voltage‑gated calcium influx and calcium‑triggered granule exocytosis as separate, linearly coupled processes. However, high‑resolution imaging shows that electrical spikes (action‑potential‑like events) and calcium waves overlap in space and time, forming a feedback loop that is highly nonlinear. To capture this, the authors propose a “spiky‑wave hybrid model”: the electrical component is described by a FitzHugh‑Nagumo two‑variable system, while the calcium dynamics follow a reaction‑diffusion equation akin to the Gray‑Scott model. The coupling term is modulated by local membrane curvature, allowing the model to reproduce experimentally observed insulin release patterns and to predict how specific genetic mutations destabilize the dynamics.
The second major case study concerns the non‑invasive manipulation of magnetic nanoparticles (MNPs) within living cells. Existing approaches often rely on Stokes‑drag or simple Markov‑type flow models, which ignore the viscoelastic nature of the cytoplasm and the dynamic reorganization of the actin network. The authors introduce a novel mathematical framework that treats the nanoparticle trajectory as a phase flow in the complex plane, while the interaction with the intracellular matrix is analyzed via nonlinear spectral methods. This approach captures the fact that the path of an MNP is continuously reshaped by the evolving cytoskeletal architecture, enabling the design of optimal magnetic field waveforms that achieve precise spatial targeting without damaging the cell.
In the discussion, the paper emphasizes that “adequate” mathematics for the cellular mesoscale must be a synthesis of several advanced fields: differential geometry to describe curved membranes, algebraic topology for organelle connectivity, and nonlinear dynamical systems for feedback‑rich biochemical networks. By aligning the mathematical language with the physical context, researchers can avoid the pitfalls of over‑simplification that plague many current models.
Finally, the authors outline future directions. They suggest that integrating high‑throughput single‑cell imaging data with the proposed topological‑dynamical framework could enable real‑time inference of cellular state and predictive control of processes such as insulin release or drug delivery via MNPs. The paper concludes that embracing Gelfand’s principle of contextual adequacy will be essential for the next generation of systems‑biology models, nanomedicine technologies, and precise intracellular manipulation strategies.