Analytical study of an exclusive genetic switch
The nonequilibrium stationary state of an exclusive genetic switch is considered. The model comprises two competing species and a single binding site which, when bound to by a protein of one species, causes the other species to be repressed. The model may be thought of as a minimal model of the power struggle between two competing parties. Exact solutions are given for the limits of vanishing binding/unbinding rates and infinite binding/unbinding rates. A mean field theory is introduced which is exact in the limit of vanishing binding/unbinding rates. The mean field theory and numerical simulations reveal that generically bistability occurs and the system is in a symmetry broken state. An exact perturbative solution which in principle allows the nonequilibrium stationary state to be computed is also developed and computed to first and second order.
💡 Research Summary
The paper presents a thorough analytical investigation of an “exclusive genetic switch,” a minimal stochastic model in which two competing protein species (or, by analogy, two competing groups) vie for a single binding site. When a protein of type 1 (X₁) occupies the site, production of type 2 proteins (X₂) is repressed, and vice‑versa; both proteins degrade at rate d, are produced at rate g when not repressed, bind at rate b, and unbind at rate u. The state of the system is fully described by the numbers of free proteins (N₁, N₂) and the binding site status S∈{0 (empty), 1 (X₁ bound), 2 (X₂ bound)}. Master equations (1‑3) capture the stochastic dynamics.
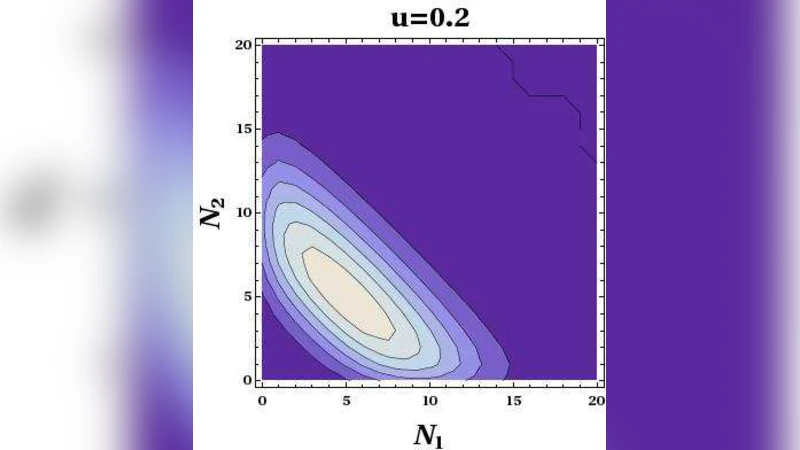
The authors first explore the model numerically using Gillespie simulations with biologically realistic parameters (g = 0.05, d = 0.005, b = 0.1, u = 0.005). The joint probability distribution P(N₁,N₂) exhibits two well‑separated peaks, each corresponding to a state where one protein dominates and the other is essentially absent—an explicit manifestation of bistability. As the unbinding rate u is increased, the two peaks move toward each other and eventually merge into a single symmetric peak, suggesting a transition from a symmetry‑broken to a symmetric regime. However, conditional distributions P₁(N₁,N₂) (when S = 1) and P₂(N₁,N₂) (when S = 2) remain asymmetric for any finite u, indicating that the apparent merging is a geometric effect rather than a genuine phase transition.
To gain analytical insight, the paper solves the model in two opposite limits. In the first limit (b, u → 0 with the binding constant k = b u held fixed), binding and unbinding are much slower than production and degradation. Consequently, the system spends long intervals in a fixed binding state, and the protein numbers follow Poisson statistics conditioned on that state. The stationary probabilities of the three binding states are derived exactly: r₀ = 1/(1 + 2k g/d), r₁ = r₂ = k g/d · r₀. When k → ∞ the symmetric state (r₀) disappears, leaving only the two asymmetric states; the switching time between them diverges, confirming strong symmetry breaking.
In the opposite limit (b, u → ∞), binding/unbinding events are fast compared to synthesis and degradation, so the binding state equilibrates instantaneously for any given (N₁,N₂). The joint probability factorizes as P(N₁,N₂) r_S(N₁,N₂) with r₀ =
Comments & Academic Discussion
Loading comments...
Leave a Comment