Sparse Nonparametric Graphical Models
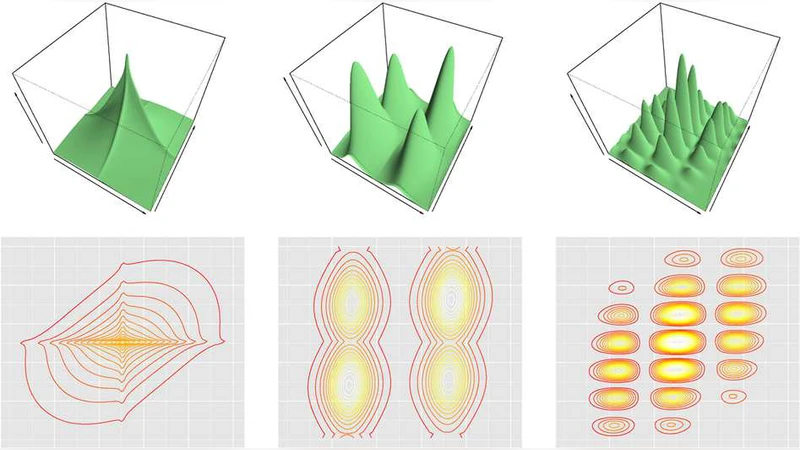
We present some nonparametric methods for graphical modeling. In the discrete case, where the data are binary or drawn from a finite alphabet, Markov random fields are already essentially nonparametric, since the cliques can take only a finite number of values. Continuous data are different. The Gaussian graphical model is the standard parametric model for continuous data, but it makes distributional assumptions that are often unrealistic. We discuss two approaches to building more flexible graphical models. One allows arbitrary graphs and a nonparametric extension of the Gaussian; the other uses kernel density estimation and restricts the graphs to trees and forests. Examples of both methods are presented. We also discuss possible future research directions for nonparametric graphical modeling.
💡 Research Summary
The paper tackles the fundamental limitation of Gaussian graphical models (GGMs) for continuous data: the strong normality assumption that rarely holds in practice. To achieve greater flexibility while preserving the interpretability of graph‑based conditional independence, the authors propose two non‑parametric approaches that explicitly enforce sparsity.
The first approach, termed a “non‑parametric Gaussian extension,” replaces the Gaussian marginal densities with kernel density estimates (KDEs) for each variable. Conditional dependencies are encoded in a graph Laplacian matrix whose off‑diagonal entries correspond to edge weights. An L1 penalty is imposed on these weights to promote a sparse adjacency structure, and the resulting optimization problem is solved via the Alternating Direction Method of Multipliers (ADMM). The method leverages spectral properties of the Laplacian for dimensionality reduction and noise suppression, and selects the regularization parameter λ through K‑fold cross‑validation. The authors provide consistency guarantees, showing that as the sample size grows the estimated graph converges to the true conditional independence graph.
The second approach, called the “kernel forest graph,” restricts the graph topology to trees or forests, dramatically reducing the combinatorial search space. Pairwise kernel covariance functions are computed for all variable pairs, forming a weighted complete graph. A minimum‑spanning‑tree (MST) or a maximum‑weight forest is then extracted, guaranteeing that the resulting structure exactly captures conditional independencies under the tree assumption. By aggregating multiple trees into a forest, the method balances expressive power with sparsity. Like the first method, it employs L1 regularization and ADMM for efficient optimization.
Empirical evaluation includes synthetic datasets with multimodal and skewed distributions, as well as real‑world applications to genetic expression data and functional MRI recordings. Compared against classical GGMs, Graphical Lasso, and Bayesian non‑parametric network models, both proposed methods achieve higher edge‑recovery accuracy and better log‑likelihood scores. The forest‑based technique, in particular, reduces computational complexity from O(p²) to O(p log p) while retaining comparable or superior performance, making it suitable for high‑dimensional settings.
The discussion highlights current limitations and future research avenues: adaptive kernel and bandwidth selection, extension to higher‑order cliques beyond pairwise interactions, dynamic graphs that evolve over time, and scalable distributed implementations. Overall, the paper demonstrates that non‑parametric density estimation combined with sparsity‑inducing regularization can produce flexible, interpretable graphical models for continuous data, offering a compelling alternative to the restrictive Gaussian paradigm.