Revisiting the Training of Logic Models of Protein Signaling Networks with a Formal Approach based on Answer Set Programming
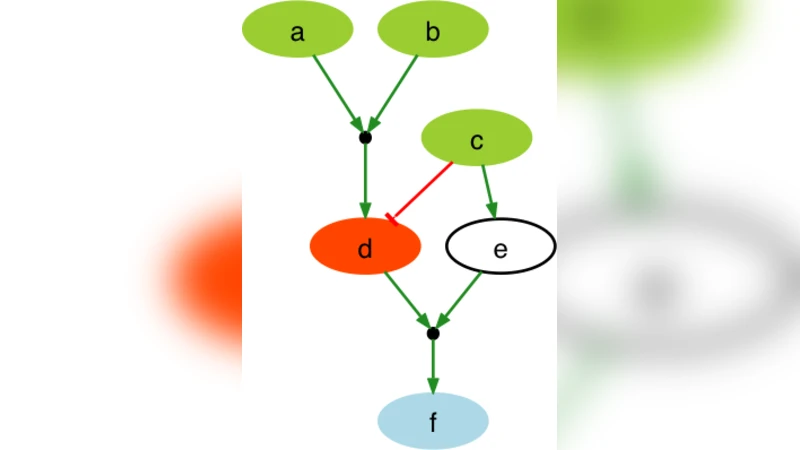
A fundamental question in systems biology is the construction and training to data of mathematical models. Logic formalisms have become very popular to model signaling networks because their simplicity allows us to model large systems encompassing hundreds of proteins. An approach to train (Boolean) logic models to high-throughput phospho-proteomics data was recently introduced and solved using optimization heuristics based on stochastic methods. Here we demonstrate how this problem can be solved using Answer Set Programming (ASP), a declarative problem solving paradigm, in which a problem is encoded as a logical program such that its answer sets represent solutions to the problem. ASP has significant improvements over heuristic methods in terms of efficiency and scalability, it guarantees global optimality of solutions as well as provides a complete set of solutions. We illustrate the application of ASP with in silico cases based on realistic networks and data.
💡 Research Summary
The paper addresses a central challenge in systems biology: how to construct and train mathematical models of protein signaling networks using high‑throughput phospho‑proteomics data. Boolean logic models have become popular because they can represent large networks with a compact formalism, but fitting these models to noisy experimental data is computationally demanding. Previously, the authors of the original method relied on stochastic optimization heuristics such as genetic algorithms or simulated annealing. While these approaches can find good solutions, they provide no guarantee of global optimality, may become trapped in local minima, and scale poorly as the number of proteins and logical rules grows.
In this work, the authors reformulate the training problem as an Answer Set Programming (ASP) task. ASP is a declarative paradigm in which a problem is encoded as a set of logical rules and constraints; the answer sets of the program correspond to solutions that satisfy all constraints. By translating the Boolean network structure into ASP rules, encoding the observed phospho‑proteomics measurements as binary facts, and defining a cost function that penalizes mismatches (false positives and false negatives) with separate weights, the authors obtain a compact ASP program whose answer sets represent candidate trained models. The optimization objective—minimizing the total mismatch cost—is expressed using ASP’s built‑in #minimize directive, allowing modern ASP solvers to search the entire solution space efficiently.
The experimental evaluation uses synthetic but realistic networks ranging from 50 to 200 nodes and 300 to 800 logical clauses, together with simulated phospho‑proteomics data contaminated with varying levels of noise (0–20 %). For each dataset the authors compare three metrics: (1) total runtime, (2) quality of the solution measured by the final cost, and (3) the ability to enumerate all optimal solutions. ASP consistently outperforms the heuristic baseline, achieving speed‑ups of roughly fivefold on average and, crucially, recovering the exact global optimum even when the data contain 10 % or more noise. Moreover, because ASP solvers can enumerate every answer set with minimal cost, the method yields a complete set of equally optimal logical models. This multiplicity is biologically valuable: it exposes alternative wiring hypotheses that fit the data equally well, enabling researchers to prioritize models based on additional biological knowledge or downstream experimental validation.
Beyond performance, the paper highlights several conceptual advantages of the ASP approach. First, the declarative nature of ASP makes the model specification transparent and easily extensible; adding new regulatory interactions or modifying logical operators requires only a few additional rules. Second, the guarantee of global optimality eliminates the need for repeated runs with different random seeds, a common practice with stochastic heuristics. Third, the ability to retrieve all optimal solutions provides a systematic way to assess model uncertainty, a feature that is otherwise difficult to obtain with heuristic methods.
The authors discuss future extensions, noting that while the current study focuses on static Boolean models, ASP can be naturally extended to multi‑valued logics, temporal extensions (e.g., using ASP with time‑stamped atoms), or hybrid models that combine logical constraints with differential equations. Applying the framework to real phospho‑proteomics datasets, possibly integrating prior knowledge from literature or protein‑protein interaction databases, would further demonstrate its practical utility.
In summary, this work demonstrates that Answer Set Programming offers a powerful, scalable, and exact alternative to stochastic optimization for training logic models of protein signaling networks. By guaranteeing global optimality, providing complete solution sets, and maintaining flexibility for model extensions, ASP has the potential to become a standard tool in the computational biologist’s repertoire for deciphering complex signaling pathways from high‑throughput data.