Evolutionary Inference for Function-valued Traits: Gaussian Process Regression on Phylogenies
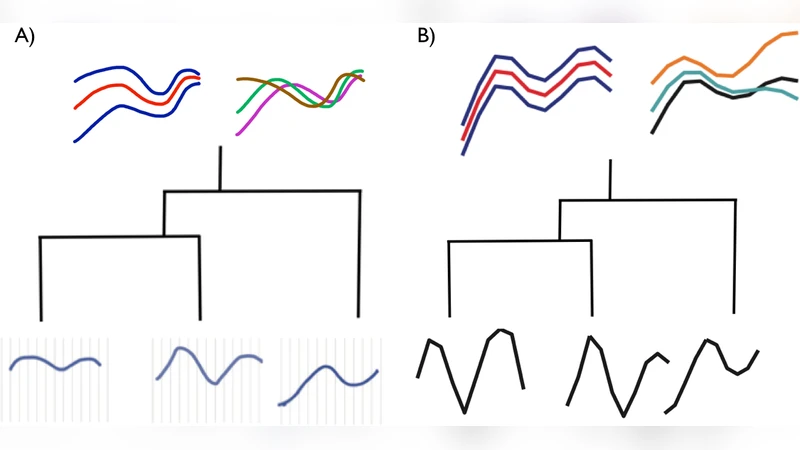
Biological data objects often have both of the following features: (i) they are functions rather than single numbers or vectors, and (ii) they are correlated due to phylogenetic relationships. In this paper we give a flexible statistical model for such data, by combining assumptions from phylogenetics with Gaussian processes. We describe its use as a nonparametric Bayesian prior distribution, both for prediction (placing posterior distributions on ancestral functions) and model selection (comparing rates of evolution across a phylogeny, or identifying the most likely phylogenies consistent with the observed data). Our work is integrative, extending the popular phylogenetic Brownian Motion and Ornstein-Uhlenbeck models to functional data and Bayesian inference, and extending Gaussian Process regression to phylogenies. We provide a brief illustration of the application of our method.
💡 Research Summary
The paper addresses a class of biological data that simultaneously exhibits two challenging properties: each observation is a continuous function rather than a scalar or vector, and observations are correlated because the species from which they are drawn share a phylogenetic history. Traditional comparative methods such as phylogenetic Brownian Motion (BM) and Ornstein‑Uhlenbeck (OU) models are designed for scalar traits and therefore cannot capture the full richness of functional data. To fill this gap, the authors propose a non‑parametric Bayesian framework that combines Gaussian Process (GP) regression with phylogenetic covariance structures.
Model construction
A Gaussian Process defines a distribution over functions. The authors extend this idea by constructing a covariance function that is the product of two components: (i) a phylogenetic kernel K_phy that depends on the evolutionary distance between two taxa (e.g., the sum of branch lengths to their most recent common ancestor) and (ii) a functional kernel K_func that measures similarity between points on the functions (e.g., squared‑exponential or Matérn). The resulting covariance between the functional value of taxon i at time t and taxon j at time t′ is
Cov
Comments & Academic Discussion
Loading comments...
Leave a Comment