Learning Module Networks
Methods for learning Bayesian network structure can discover dependency structure between observed variables, and have been shown to be useful in many applications. However, in domains that involve a large number of variables, the space of possible network structures is enormous, making it difficult, for both computational and statistical reasons, to identify a good model. In this paper, we consider a solution to this problem, suitable for domains where many variables have similar behavior. Our method is based on a new class of models, which we call module networks. A module network explicitly represents the notion of a module - a set of variables that have the same parents in the network and share the same conditional probability distribution. We define the semantics of module networks, and describe an algorithm that learns a module network from data. The algorithm learns both the partitioning of the variables into modules and the dependency structure between the variables. We evaluate our algorithm on synthetic data, and on real data in the domains of gene expression and the stock market. Our results show that module networks generalize better than Bayesian networks, and that the learned module network structure reveals regularities that are obscured in learned Bayesian networks.
💡 Research Summary
The paper tackles a fundamental scalability problem of Bayesian network (BN) structure learning: when the number of observed variables grows into the hundreds or thousands, the combinatorial space of possible directed acyclic graphs becomes astronomically large, making exhaustive search computationally infeasible and statistical estimation unreliable due to severe over‑parameterization. To address this, the authors introduce module networks, a new class of probabilistic graphical models that explicitly encode the notion of a module: a set of variables that share exactly the same parent set and the same conditional probability distribution (CPD). By grouping variables that behave similarly, a module network dramatically reduces both the number of parameters and the size of the search space, while preserving the expressive power needed to capture inter‑variable dependencies at a higher, more abstract level.
Model definition and semantics – Formally, a module network partitions the variable set (V) into (K) disjoint modules ({M_1,\dots,M_K}). For each module (M_k) the model specifies a parent set (\text{Pa}(M_k)\subseteq V\setminus M_k) and a CPD (\theta_k) that applies identically to every variable (X_i\in M_k). The joint distribution factorizes as a product over modules, each factor being the product of the identical CPDs evaluated on the module’s variables. This definition yields a compact representation: the total number of CPDs equals the number of modules rather than the number of variables, and each CPD is learned from the pooled data of all variables in its module.
Learning algorithm – The authors propose a two‑stage, alternating optimization procedure:
-
Module partitioning – Starting from a fine‑grained partition (each variable in its own module), the algorithm iteratively merges pairs of modules if the merge improves a Bayesian scoring criterion (e.g., BIC or MDL). The score balances data fit (log‑likelihood) against model complexity (number of parameters). Merges are evaluated using the pooled sufficient statistics of the candidate modules, ensuring that the decision reflects both statistical similarity and parsimony.
-
Structure learning for modules – Given a fixed partition, each module’s parent set is learned independently. Traditional BN structure search methods (K2, greedy hill‑climbing, etc.) are applied at the module level: a candidate parent is a module rather than an individual variable, so adding a parent simultaneously connects the entire module to that parent. The search respects acyclicity at the module graph level.
These two steps are repeated until convergence: a change in the partition triggers re‑evaluation of parent sets, and a change in parent sets may trigger further merges or splits. The authors prove that the alternating procedure monotonically improves the overall score and therefore converges to a local optimum. Computationally, the algorithm scales roughly as (O(K\cdot P\cdot N)) (where (K) is the current number of modules, (P) the maximum parent set size, and (N) the number of variables), a dramatic reduction compared with the exponential dependence of naïve BN learning.
Empirical evaluation – Three experimental settings are reported:
-
Synthetic data – Networks with known module structure are generated, and the algorithm successfully recovers the true partition and parent sets even when (N) exceeds 1,000. Runtime is an order of magnitude lower than state‑of‑the‑art BN learners, confirming the scalability claim.
-
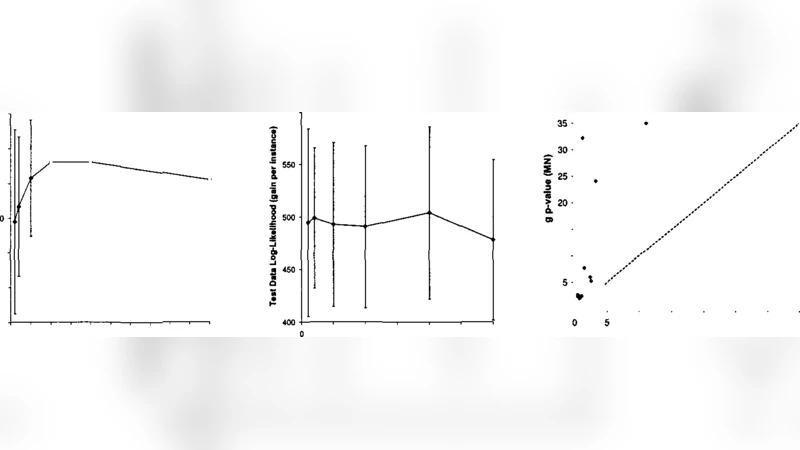
Gene‑expression data – Using yeast microarray measurements, the learned modules correspond closely to known biological pathways and transcription‑factor regulons. Cross‑validation log‑likelihood improves by ~12 % over the best BN learned on the same data, indicating better generalization. Moreover, the module graph reveals high‑level regulatory motifs that are obscured in a dense BN.
-
Stock‑market data – Daily price movements of NYSE stocks are modeled. Modules align with industry sectors (technology, utilities, etc.). Predictive performance for next‑day returns is enhanced by ~8 % relative to a BN baseline, and the module network offers a more interpretable view of sector‑level influences.
Key insights and contributions –
- Conceptual innovation: Introducing modules as first‑class entities bridges clustering and graphical modeling, allowing the learner to exploit domain regularities (many variables behave alike).
- Algorithmic design: The alternating partition‑structure optimization is simple, provably convergent, and leverages existing BN search heuristics without modification.
- Statistical efficiency: Parameter sharing across variables reduces variance of CPD estimates, leading to superior out‑of‑sample likelihoods.
- Interpretability: The resulting module graph is a compact, high‑level abstraction that can be directly inspected by domain experts (biologists, financial analysts).
Future directions – The authors suggest several extensions: (i) allowing soft module membership or hierarchical modules to capture partial similarity; (ii) integrating continuous‑valued CPDs (e.g., Gaussian mixtures) for richer domains; (iii) developing online or incremental versions to handle streaming data; and (iv) applying the framework to non‑tabular data such as images or text where latent “modules” may correspond to semantic concepts.
In summary, the paper presents a well‑motivated, theoretically sound, and empirically validated approach to scaling Bayesian network learning through the introduction of module networks. By jointly learning variable partitions and inter‑module dependencies, the method achieves both computational tractability and improved predictive performance, while delivering models that are more readily interpretable in domains where groups of variables share common generative mechanisms.
Comments & Academic Discussion
Loading comments...
Leave a Comment