Molecular dynamics simulation of reversibly self-assembling shells in solution using trapezoidal particles
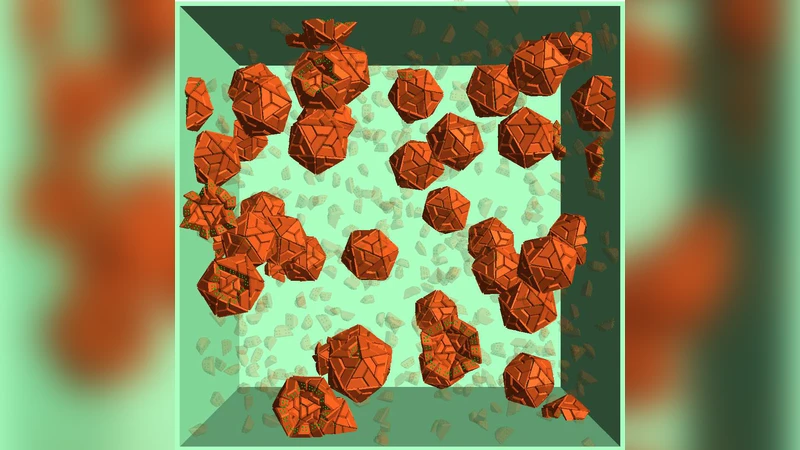
The self-assembly of polyhedral shells, each constructed from 60 trapezoidal particles, is simulated using molecular dynamics. The spatial organization of the component particles in this shell is similar to the capsomer proteins forming the capsid of a T=1 virus. Growth occurs in the presence of an atomistic solvent and, under suitable conditions, achieves a high yield of complete shells. The simulations provide details of the structure and lifetime of the particle clusters that appear as intermediate states along the growth pathway, and the nature of the transitions between them. In certain respects the growth of size-60 shells from trapezoidal particles resembles the growth of icosahedral shells from triangular particles studied previously, with reversible bonding playing a major role in avoiding incorrect assembly, although the details differ due to particle shape and bond organization. The strong preference for maximal bonding exhibited by the triangular particle clusters is also apparent for trapezoidal particles, but this is now confined to early growth, and is less pronounced as shells approach completion along a variety of pathways.
💡 Research Summary
This paper presents a comprehensive molecular‑dynamics (MD) study of the reversible self‑assembly of polyhedral shells composed of sixty trapezoidal particles, a geometry that mirrors the capsomers of a T = 1 viral capsid. Each particle is a five‑face, non‑symmetric trapezoid whose binding sites are placed on specific faces; the interactions are modeled with distance‑ and angle‑dependent potentials that emulate protein‑protein contacts. An explicit atomistic solvent is included, allowing realistic friction, collisions, and rotational diffusion of the particles. By systematically varying temperature (expressed as ε/kBT) and particle concentration, the authors identify a narrow thermodynamic window (ε/kBT ≈ 0.5–0.7) in which the system yields a high proportion of complete shells—approximately 85 % of the particles end up in correctly assembled 60‑particle shells.
The assembly pathway is dissected through cluster‑size distributions, lifetimes, and structural analyses. Early intermediates (5–10 particles) display a strong preference for maximal bonding; these clusters are long‑lived because each particle can form up to four or five bonds, creating a deep energetic well that stabilizes the nucleus. As growth proceeds to 20–40 particles, multiple pathways emerge: some clusters continue to add particles in a highly ordered fashion, while others become trapped in partially bonded configurations that persist for extended periods. This diversification is a direct consequence of the trapezoidal shape, which provides more binding orientations than the triangular particles studied previously.
Reversibility of the bonds is a central theme. Erroneous contacts are broken by thermal fluctuations, and the solvent‑mediated collisions enable the particles to re‑orient and re‑bind correctly. This “error‑correction” mechanism mirrors the kinetic proofreading observed in real viral capsid assembly and is essential for avoiding kinetic traps that would otherwise dominate the landscape. Notably, while triangular particles exhibit a global bias toward maximal bonding throughout the entire assembly, trapezoidal particles retain this bias only in the early stages; near completion (≈45–60 particles) the system tolerates a variety of bonding sequences, yet still converges on the correct shell because the reversible dynamics continually prune defective configurations.
The authors compare their findings with earlier work on icosahedral shells built from triangular units. Both systems rely on reversible bonding to achieve high yields, but the trapezoidal model demonstrates that particle shape and bond topology can shift the balance between deterministic nucleation and stochastic pathway branching. The study therefore provides valuable design principles for synthetic nanostructures: by tuning particle geometry, interaction specificity, and solvent conditions, one can steer the self‑assembly process toward desired architectures with high fidelity. The implications extend to virus‑like particle engineering, drug‑delivery capsule fabrication, and the broader field of programmable matter.
Comments & Academic Discussion
Loading comments...
Leave a Comment