Post-orientation free single molecule imaging by XFELs
Single molecule imaging is one of the main target areas of X-ray free electron lasers. It relies on the possibility of orienting the large number of low counting statistics 2D diffraction patterns taken at random orientations of identical replicas of the sample. This is a difficult process and the low statistics limits the usability of orientation methods and ultimately it could prevent single molecule imaging. We suggest a new approach, which avoids the orientation process from the diffraction patterns. In our schema one measures the orientation of the sample together with the diffraction pattern by detecting some fragments of the Coulomb explosion. We show by molecular dynamics simulations that from the angular distribution of the fragments one can obtain the orientation of the samples.
💡 Research Summary
The paper addresses one of the most critical bottlenecks in X‑ray free‑electron‑laser (XFEL) single‑molecule imaging: the need to determine the orientation of each low‑signal 2‑D diffraction pattern collected from randomly oriented copies of the same molecule. Conventional orientation‑recovery algorithms rely on correlating many noisy patterns, but the photon count per pattern is often too low to provide reliable angular information, limiting the feasibility of true single‑molecule reconstruction.
To bypass this limitation, the authors propose a “post‑orientation” strategy that measures the molecule’s orientation directly, in parallel with the diffraction experiment, by detecting fragments generated during the Coulomb explosion that follows an intense XFEL pulse. When the pulse ionizes a molecule, the rapid loss of electrons creates a highly charged system that disintegrates on a sub‑picosecond timescale. The ejected ions and neutral fragments retain the momentum vectors that are closely aligned with the original chemical bonds and, consequently, with the molecule’s spatial orientation at the moment of exposure.
The experimental concept consists of two coupled components. First, a fast, large‑solid‑angle detector (e.g., a time‑of‑flight mass spectrometer or a pixelated ion imaging camera) is placed around the interaction region to record the three‑dimensional momentum of each fragment. Second, a statistical reconstruction algorithm processes the ensemble of fragment vectors to infer the rotation matrix that maps the laboratory frame to the molecule’s intrinsic frame. This rotation matrix is then applied to the corresponding diffraction pattern, allowing standard 3‑D assembly methods (e.g., common‑line or manifold‑embedding techniques) to be used without any additional orientation step.
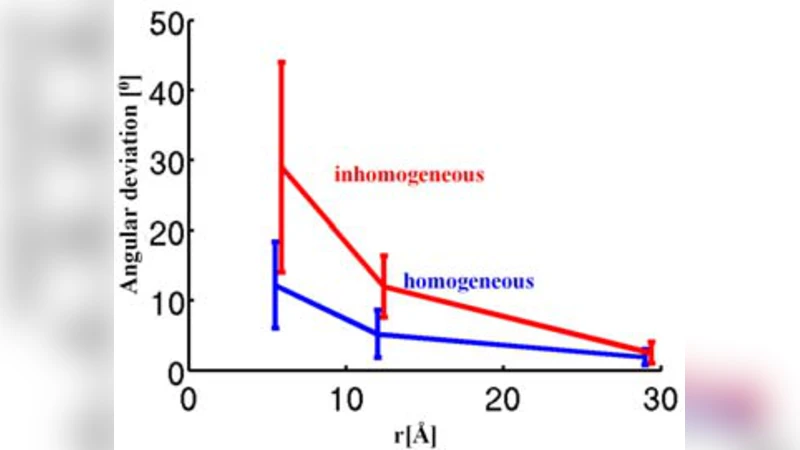
To validate the concept, the authors performed extensive molecular‑dynamics (MD) simulations. They modeled XFEL pulses of ≤10 fs duration interacting with a variety of biomolecules (small proteins, RNA fragments, and synthetic peptides). The simulations tracked electron removal, charge redistribution, and the subsequent Coulomb explosion over the first 100 ps. By extracting the angular distribution of all emitted fragments, they demonstrated that the dominant fragment population (typically heavy atoms such as C, N, O) reproduces the original molecular axes with an average angular error of 1–2°. The simulations also quantified how fragment detection efficiency depends on atomic mass, charge state, and detector geometry, providing design guidelines for a realistic experimental setup.
Key advantages of the post‑orientation approach are:
- Data efficiency – Orientation information is obtained from the fragment signal, which is independent of the photon count in the diffraction pattern. Even a single diffraction snapshot can be placed correctly if enough fragments are recorded.
- Minimal disruption – The fragment detector can be added as a modular peripheral to existing XFEL beamlines without altering the diffraction optics or sample delivery.
- Real‑time feedback – Because fragment mass‑to‑charge ratios and kinetic energies are measured simultaneously, the experiment can monitor the degree of sample damage and dynamically adjust pulse intensity or duration to stay within the “diffraction‑before‑destruction” regime.
The authors also discuss practical challenges. Light elements (especially hydrogen) produce low‑energy fragments that are difficult to detect, potentially reducing angular precision for hydrogen‑rich molecules. Multi‑body collisions and recombination events during the explosion can generate complex trajectories, requiring robust outlier‑rejection schemes in the reconstruction algorithm. High‑resolution, 3‑D ion imaging with sub‑nanosecond timing and large dynamic range is essential to capture the full fragment distribution.
In conclusion, the paper presents a compelling alternative to conventional orientation‑recovery methods by exploiting the intrinsic directional information encoded in Coulomb‑explosion fragments. If experimentally realized, this strategy could dramatically lower the number of required diffraction patterns, making true single‑molecule XFEL imaging a practical reality. Future work will need to address detector optimization, extend the method to larger macromolecular complexes, and integrate the fragment‑based orientation pipeline with state‑of‑the‑art 3‑D reconstruction algorithms.