An experimental evaluation of de-identification tools for electronic health records
The robust development of Electronic Health Records (EHRs) causes a significant growth in sharing EHRs for clinical research. However, such a sharing makes it difficult to protect patient’s privacy. A number of automated de-identification tools have been developed to reduce the re-identification risk of published data, while preserving its statistical meaning. In this paper, we focus on the experimental evaluation of existing automated de-identification tools, as applied to our EHR database, to assess which tool performs better with each quasi-identifiers defined in our paper. Performance of each tool is analyzed wrt. two aspects: individual disclosure risk and information loss. Through this experiment, the generalization method has better performance on reducing risk and lower degree of information loss than suppression, which validates it as more appropriate de-identification technique for EHR databases.
💡 Research Summary
The paper presents a systematic experimental evaluation of several automated de‑identification tools applied to a real electronic health record (EHR) database. Recognizing that the rapid expansion of EHR sharing for clinical research heightens the risk of patient re‑identification, the authors aim to determine which tool and which technique (generalization versus suppression) best balance privacy protection with data utility.
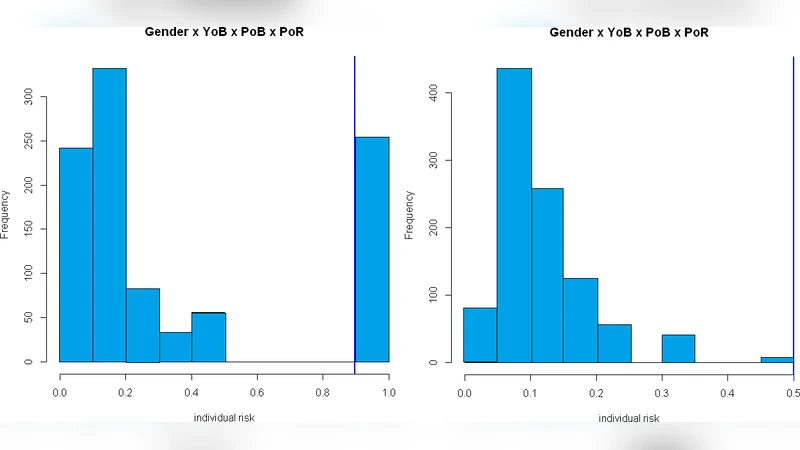
First, the authors define a set of quasi‑identifiers (QIs) that are common in their EHR repository, including age, gender, admission date, department, diagnosis codes, and prescription codes. They construct a risk model based on k‑anonymity, measuring the probability that an individual record can be uniquely linked to an external data source given a particular combination of QIs. Two quantitative performance metrics are introduced: (1) individual disclosure risk, which estimates the likelihood of successful re‑identification for each record, and (2) information loss, which quantifies the statistical deviation between the original and de‑identified datasets (distributional changes, correlation preservation, entropy shift).
Four de‑identification tools are selected for comparison: two commercial solutions (e.g., ARX) and two open‑source frameworks (e.g., sdcMicro, a privacy‑preserving data publishing toolkit). Each tool supports both generalization (e.g., converting exact ages to 5‑year intervals, aggregating dates to month level) and suppression (e.g., replacing values with asterisks or nulls, removing entire attributes). The experiments are conducted under identical QI sets and three k‑values (k = 5, 10, 20) to assess scalability of privacy guarantees.
Results consistently show that generalization outperforms suppression across all tools. Generalization reduces individual disclosure risk to levels comparable to or lower than suppression while incurring substantially less information loss. This advantage is most pronounced for continuous attributes such as age and admission date, where interval‑based generalization preserves mean, variance, and overall distribution. Suppression, by contrast, eliminates or masks entire records, leading to a noticeable reduction in sample size, especially for rare disease cohorts, and consequently degrading statistical power.
Tool‑specific differences also emerge. Although all tools implement the same high‑level algorithms, variations in optimization strategies, automatic parameter tuning, and support for user‑defined generalization hierarchies affect the final risk‑utility trade‑off. For instance, ARX’s multi‑objective optimization seeks the minimal information loss solution for a given k, whereas sdcMicro’s default settings produce coarser generalizations and higher information loss.
Based on these findings, the authors recommend the following practical guidelines for EHR de‑identification:
- Prefer generalization when the primary analytical goals involve population‑level statistics or trend analysis, as it maintains data fidelity while achieving required privacy levels.
- For studies focusing on small, high‑risk subpopulations (e.g., rare disease research), combine fine‑grained generalization with additional privacy mechanisms such as differential privacy rather than relying on blunt suppression.
- Evaluate de‑identification tools not only on their privacy guarantees but also on implementation efficiency, ease of parameter configuration, and flexibility for custom hierarchies.
- Conduct post‑de‑identification audits, including re‑identification risk assessments and utility checks, to ensure an ongoing balance between privacy and data usefulness.
In conclusion, the experimental comparison validates that, for the authors’ EHR dataset, generalization offers a superior balance of reduced re‑identification risk and lower information loss compared to suppression. This evidence supports the adoption of generalization‑centric de‑identification strategies in health informatics platforms and informs policy decisions regarding the safe sharing of clinical data.
Comments & Academic Discussion
Loading comments...
Leave a Comment