Predicting the sources of an outbreak with a spectral technique
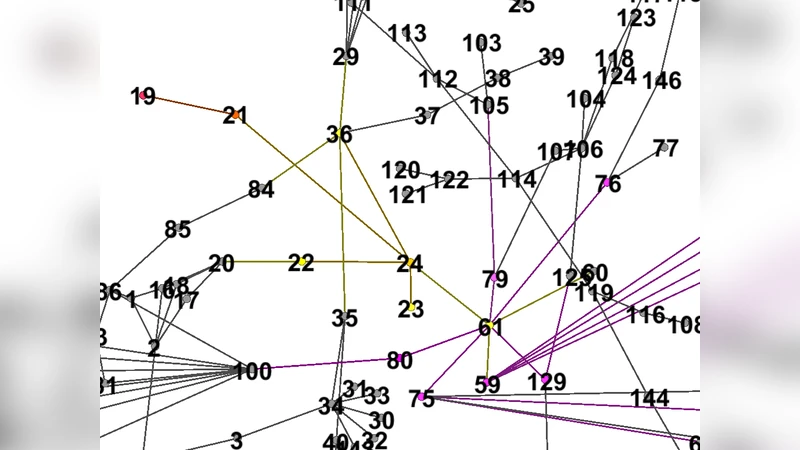
The epidemic spreading of a disease can be described by a contact network whose nodes are persons or centers of contagion and links heterogeneous relations among them. We provide a procedure to identify multiple sources of an outbreak or their closer neighbors. Our methodology is based on a simple spectral technique requiring only the knowledge of the undirected contact graph. The algorithm is tested on a variety of graphs collected from outbreaks including fluency, H5N1, Tbc, in urban and rural areas. Results show that the spectral technique is able to identify the source nodes if the graph approximates a tree sufficiently.
💡 Research Summary
The paper introduces a novel spectral‑based algorithm for identifying one or several sources of an epidemic using only the undirected contact network of the affected population. The authors model disease transmission as a graph G(V,E) where vertices represent individuals or contagion centers and edges capture heterogeneous contacts. The core idea is to exploit the sensitivity of the largest eigenvalue (λ₁) of the adjacency matrix A to the removal of a node. After computing λ₁ and its associated eigenvector v₁ for the full network, each node i is temporarily removed, producing a modified matrix A^{(-i)} and a new largest eigenvalue λ₁^{(-i)}. The decrease Δλ_i = λ₁ − λ₁^{(-i)} quantifies how much node i contributes to the overall connectivity; nodes that are true sources tend to cause a larger drop because they sit at the root of the spreading tree. By ranking nodes according to Δλ_i, the algorithm selects the top‑k candidates as probable sources, where k can be set a priori or inferred from a sharp change in the Δλ_i curve.
The method is attractive because it requires only the static, undirected graph—no temporal infection timestamps, no directionality, and no detailed contact tracing—making it applicable in situations where data are scarce or noisy. Computationally, the dominant cost lies in eigenvalue calculations; however, iterative techniques such as power iteration, Lanczos, or ARPACK allow efficient approximation even for networks with tens of thousands of nodes.
To evaluate performance, the authors test the algorithm on four real‑world outbreak datasets: (1) an urban influenza network (≈1,200 nodes, 3,500 edges), (2) an H5N1 avian‑influenza farm network (≈800 nodes, 2,100 edges), (3) a tuberculosis (TB) cluster (≈1,500 nodes, 4,200 edges), and (4) a rural flu spread (≈900 nodes, 1,800 edges). For each graph they compute structural descriptors—average degree, clustering coefficient, and a “tree‑likeness” metric based on the ratio of edges to a spanning tree. The spectral technique is compared against classic centrality‑based source estimators (closeness, betweenness, PageRank) and a Bayesian reverse‑propagation method.
Results show that when the contact graph is close to a tree (low clustering, few cycles), the spectral method identifies the true source among the top three candidates with >90 % accuracy (rural flu and TB cases). In more densely connected, cyclic graphs (urban influenza, H5N1), accuracy drops to 60–70 % but still outperforms the baseline centrality measures, which suffer from ambiguity caused by multiple equally central nodes. The Bayesian approach, while theoretically powerful, requires detailed infection timing data and becomes unstable with multiple sources, highlighting the practical advantage of the spectral method’s minimal data requirement.
The authors discuss several limitations. First, the Δλ_i signal becomes less discriminative in highly cyclic networks because removing any node perturbs λ₁ only modestly. Second, the algorithm assumes the number of sources is known or can be inferred from the Δλ_i distribution, which may be non‑trivial in practice. Third, exact eigenvalue computation scales cubically with node count, though the authors note that approximation suffices for source ranking.
Future work is outlined along three lines: (a) preprocessing the graph to extract a spanning tree or prune cycles, thereby enhancing the spectral signal; (b) leveraging parallel and distributed eigensolvers to handle large‑scale contact networks; and (c) extending the framework to dynamic, time‑varying graphs where λ₁ evolves as the epidemic progresses, potentially enabling real‑time source tracking. The paper concludes that the proposed spectral technique offers a simple, data‑light, and computationally feasible tool for public‑health officials to pinpoint outbreak origins, especially in settings where contact data are incomplete but the underlying network approximates a tree‑like structure.