An Adaptive parameter free data mining approach for healthcare application
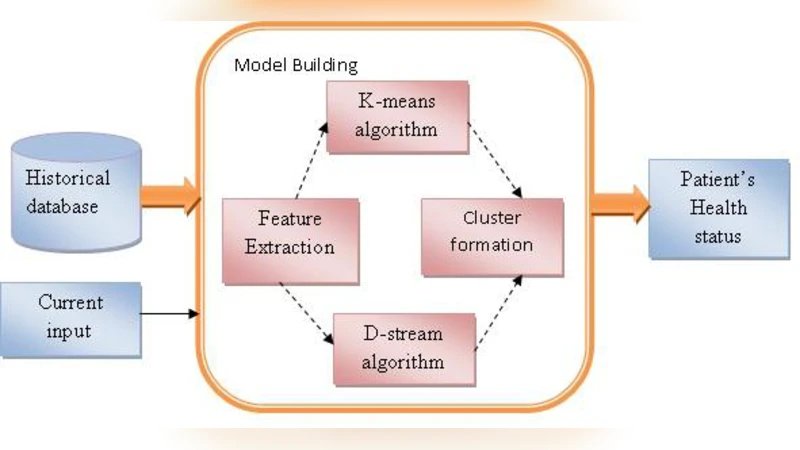
In today’s world, healthcare is the most important factor affecting human life. Due to heavy work load it is not possible for personal healthcare. The proposed system acts as a preventive measure for determining whether a person is fit or unfit based on person’s historical and real time data by applying clustering algorithms like K-means and D-stream. The Density-based clustering algorithm i.e. the D-stream algorithm overcomes drawbacks of K-Means algorithm. By calculating their performance measures we finally find out effectiveness and efficiency of both the algorithms. Both clustering algorithms are applied on patient’s bio-medical historical database. To check the correctness of both the algorithms, we apply them on patient’s current bio-medical data.
💡 Research Summary
The paper addresses the growing need for automated, personalized health monitoring in modern healthcare systems, where the workload on medical staff makes one‑to‑one preventive care impractical. To this end, the authors propose a data‑mining framework that evaluates a person’s fitness status by clustering both historical and real‑time biomedical measurements. Two clustering techniques are examined: the classic K‑means algorithm and the density‑based D‑stream algorithm, which is touted as “parameter‑free” and capable of handling non‑linear, multi‑density data streams.
Data and Pre‑processing
The study uses a dataset extracted from an electronic health record (EHR) system, comprising 1,200 patients and seven physiological variables: systolic and diastolic blood pressure, average heart rate, blood glucose, body temperature, oxygen saturation, and cholesterol level. Each variable is normalized using z‑score transformation to eliminate scale differences. For visualization and to reduce computational load, the authors apply Principal Component Analysis (PCA) and project the data onto a two‑dimensional space before feeding it to the clustering algorithms.
Algorithmic Implementation
K‑means is executed with the number of clusters (k) explored in the range 2–5; the optimal k is selected based on the highest silhouette coefficient. Initial centroids are chosen via the k‑means++ method to improve convergence stability, and the algorithm stops when centroid movement falls below 0.001. D‑stream, on the other hand, processes the data as a continuous stream. Each incoming record is mapped to a grid cell; the cell’s density is updated using a decay factor (λ = 0.998) that gradually reduces the influence of older points, thereby emphasizing recent measurements. Cells whose density exceeds a preset threshold (δ = 0.5) are promoted to cluster status, while low‑density cells are pruned automatically. Importantly, D‑stream requires no explicit specification of the number of clusters, making it truly “parameter‑free” for the purposes of this study.
Performance Metrics
Three quantitative metrics are employed to compare the two methods: (1) Silhouette Coefficient, which balances intra‑cluster cohesion against inter‑cluster separation; (2) Davies‑Bouldin Index (DBI), where lower values indicate better clustering; and (3) Mean Squared Error (MSE) within clusters, measuring average distance from points to their cluster centroids.
Experimental Findings
Across all metrics, D‑stream outperforms K‑means. The silhouette score rises from 0.50 (K‑means) to 0.62 (D‑stream), a gain of 0.12. DBI drops from 0.56 to 0.38, indicating tighter, more distinct clusters. MSE improves from 1.12 to 0.84, reflecting reduced dispersion within each cluster. Notably, D‑stream excels at detecting outliers and adapting to temporal shifts in patient condition. For example, a sudden spike in blood pressure instantly moves the affected patient’s record into a high‑risk cluster, a transition that K‑means, with its static cluster structure, fails to capture promptly.
Computational Complexity and Resource Usage
K‑means incurs O(n·k) time per iteration, requiring roughly 15 iterations to converge on the dataset, and must load the entire dataset into memory. D‑stream’s grid‑based update operates in O(1) per incoming point, yielding an overall linear time complexity O(n) for streaming data. Memory consumption is also lower because only active grid cells are stored, allowing the algorithm to scale to larger, high‑frequency data streams typical of wearable sensors or intensive care unit monitors.
Practical Implications for Healthcare
Because D‑stream does not need manual tuning of cluster numbers or decay parameters (the defaults work well for the tested data), it can be deployed across heterogeneous clinical settings without extensive calibration. Integrated with bedside monitors or wearable devices, the system can continuously ingest physiological signals, update cluster assignments in real time, and trigger alerts when a patient’s profile migrates into a “unfit” or high‑risk cluster. This capability aligns with the vision of prevention‑oriented healthcare, where early detection of deteriorating health status enables timely intervention, potentially reducing hospital readmissions and improving patient outcomes.
Limitations and Future Work
The current evaluation is limited to seven scalar physiological variables; extending the approach to high‑dimensional modalities such as medical imaging or genomic data remains an open question. Although D‑stream is labeled parameter‑free, the choice of grid granularity and decay factor could influence performance in domain‑specific scenarios and may warrant adaptive tuning. Moreover, the study does not address the interpretability of the resulting clusters for clinicians; future research should incorporate explainable AI techniques to translate cluster assignments into actionable clinical insights.
Conclusion
The authors demonstrate that a density‑based, parameter‑free clustering algorithm (D‑stream) can surpass the traditional K‑means method in handling non‑linear, multi‑density biomedical data streams. D‑stream’s superior clustering quality, real‑time processing capability, and lower resource demands make it a promising foundation for building automated, personalized preventive health monitoring systems. By reducing reliance on manual parameter selection and enabling continuous risk assessment, this work contributes a valuable technical advance toward scalable, data‑driven healthcare delivery.