Field theory approach in the dynamics of biomatter
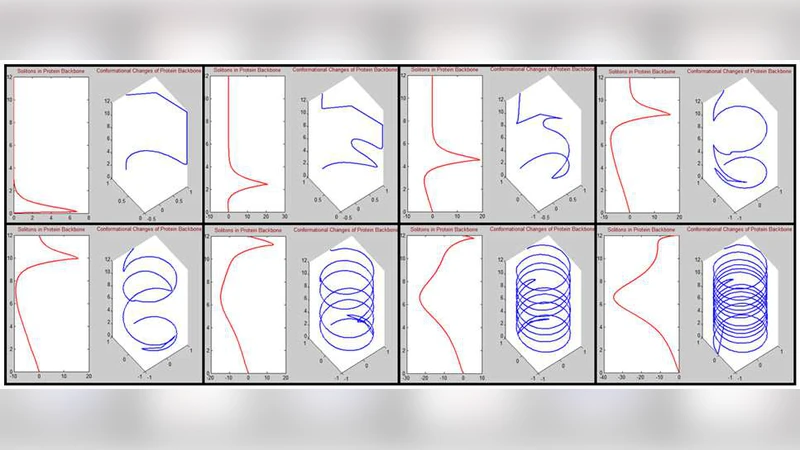
A new approach to model the biomatter dynamics based on the field theory is presented. It is shown that some well known tools in field theory can be utilized to describe the physical phenomena in life matters, in particular at elementary biomatters like DNA and proteins. In this approach, the biomatter dynamics are represented as results of interactions among its elementary matters in the form of lagrangian. Starting from the lagrangian would provide stronger underlying theoretical consideration for further extension. Moreover, it also enables us to acquire rich physical observables using statistical mechanics instead of relying on the space-time dynamics from certain equation of motions which is not solvable due to its nonlinearities. Few examples from previous results are given and explained briefly.
💡 Research Summary
The paper proposes a novel theoretical framework for describing the dynamics of biomatter—specifically elementary biological entities such as DNA and proteins—by importing the machinery of quantum field theory (QFT) into the realm of biophysics. Rather than relying on conventional molecular dynamics or stochastic differential equations, which quickly become intractable due to strong nonlinearities, the authors start from a Lagrangian density that encodes the kinetic, mass, and interaction terms of the relevant fields. In this picture, a DNA double helix is represented by two coupled scalar fields ϕ₁(x, t) and ϕ₂(x, t), while a protein chain is modeled as a multi‑component vector or spinor field ψᵢ(x, t) living in an internal symmetry space (e.g., SU(N)).
The total Lagrangian is split into a free part L₀, containing the usual kinetic (∂_μϕ∂^μϕ) and mass (m²ϕ²) contributions, and an interaction part L_int, which can be expressed either as a self‑interaction potential V(ϕ, ψ) (for example a ϕ⁴ term λϕ⁴/4! that captures nonlinear elasticity and electrostatic effects in DNA) or as a gauge‑covariant derivative term D_μϕ D^μϕ that introduces minimal coupling to an auxiliary gauge field A_μ. By imposing symmetries—U(1) for charge conservation in the DNA sector, SU(N) for the internal degrees of freedom of proteins—the model inherits conserved currents and Ward identities that greatly simplify perturbative expansions and renormalization.
A key advantage of the Lagrangian approach is that it provides a natural bridge to statistical mechanics via the path‑integral formalism. The authors define the partition function
Z = ∫𝔻ϕ 𝔻ψ exp
Comments & Academic Discussion
Loading comments...
Leave a Comment