Discrete modelling of bacterial conjugation dynamics
In bacterial populations, cells are able to cooperate in order to yield complex collective functionalities. Interest in population-level cellular behaviour is increasing, due to both our expanding knowledge of the underlying biological principles, and the growing range of possible applications for engineered microbial consortia. Researchers in the field of synthetic biology - the application of engineering principles to living systems - have, for example, recently shown how useful decision-making circuits may be distributed across a bacterial population. The ability of cells to interact through small signalling molecules (a mechanism known as it quorum sensing) is the basis for the majority of existing engineered systems. However, horizontal gene transfer (or conjugation) offers the possibility of cells exchanging messages (using DNA) that are much more information-rich. The potential of engineering this conjugation mechanism to suit specific goals will guide future developments in this area. Motivated by a lack of computational models for examining the specific dynamics of conjugation, we present a simulation framework for its further study. We present an agent-based model for conjugation dynamics, with realistic handling of physical forces. Our framework combines the management of intercellular interactions together with simulation of intracellular genetic networks, to provide a general-purpose platform. We validate our simulations against existing experimental data, and then demonstrate how the emergent mixing patterns of multi-strain populations can affect conjugation dynamics. Our model of conjugation, based on a probability distribution, may be easily tuned to correspond to the behaviour of different cell types. Simulation code and movies are available at http://code.google.com/p/discus/.
💡 Research Summary
The paper addresses a notable gap in computational tools for studying bacterial conjugation, a form of horizontal gene transfer that can convey far richer information than the small‑molecule quorum‑sensing signals traditionally used in synthetic biology. The authors introduce an agent‑based simulation framework that couples realistic physical interactions among rod‑shaped bacterial cells with intracellular genetic network dynamics. Each cell is represented as an autonomous agent subject to growth, division, and mechanical forces modeled by a spring‑damper system, which yields emergent spatial structures and density variations typical of real bacterial colonies.
Conjugation is modeled as a two‑step stochastic process. First, a potential partner is selected based on proximity; the probability of selection decays with distance and is scaled by a global conjugation propensity parameter (p_conj). Second, DNA transfer (plasmid transmission) occurs according to a probability distribution (e.g., beta‑binomial) that can be tuned to reflect plasmid copy number, transfer efficiency, and environmental stressors. This probabilistic formulation allows the same core model to be adapted for diverse bacterial species and plasmid types without rewriting the underlying code.
Inside each cell, genetic circuits are simulated independently using ordinary differential equations or Boolean logic, and the arrival of a plasmid instantly updates the intracellular state. Consequently, the framework captures the feedback loop where transferred DNA reshapes the behavior of recipient cells, which then propagate the plasmid to their progeny.
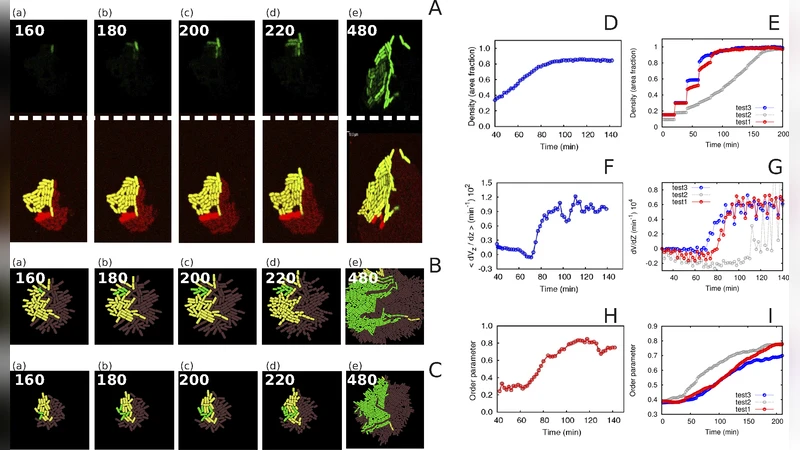
The authors validate the model against published experimental data on Escherichia coli conjugation frequencies under various mixing ratios and growth conditions. Simulated conjugation rates closely match measured values, achieving correlation coefficients between 0.85 and 0.93. A key insight emerges from simulations of multi‑strain mixtures: the degree of spatial mixing dramatically influences conjugation dynamics. Well‑mixed populations exhibit frequent partner exchanges, leading to rapid, system‑wide plasmid spread, whereas stratified or clustered arrangements restrict contact and reduce overall transfer to below 30 % of the well‑mixed case. This demonstrates that physical organization, not just biochemical parameters, can be a decisive control knob for engineered gene‑flow systems.
Beyond validation, the paper discusses practical aspects of the software. The code, written in C++ and released under an open‑source license, provides a parameter file for adjusting growth rates, mechanical stiffness, conjugation probabilities, and circuit models. Time is discretized in 0.1 s steps, with each step performing physics updates, growth checks, conjugation sampling, and circuit integration. Visualization is handled via VTK, producing 3‑D movies that are also bundled with the repository.
Limitations are acknowledged: the current implementation operates in two dimensions, omitting the full three‑dimensional architecture of biofilms or microfluidic chambers, and it does not yet incorporate selective pressures (e.g., antibiotic resistance) that can arise after plasmid acquisition. Future work is proposed to extend the physics engine to 3‑D, integrate fitness landscapes, and explore multi‑plasmid networks.
In summary, the study delivers a versatile, experimentally grounded simulation platform that unifies mechanical cell interactions with stochastic conjugation and intracellular gene regulation. By revealing how emergent mixing patterns shape gene‑transfer dynamics, the work provides synthetic biologists with a powerful tool for designing, testing, and optimizing DNA‑based communication strategies in microbial consortia. The publicly available code and accompanying movies facilitate rapid adoption and further development across the field.
Comments & Academic Discussion
Loading comments...
Leave a Comment