Dynamic Landscapes: A Model of Context and Contingency in Evolution
The basic mechanics of evolution have been understood since Darwin. But debate continues over whether macroevolutionary phenomena are driven primary by the fitness structure of genotype space or by ecological interaction. In this paper we propose a simple, abstract model capturing some key features of fitness-landscape and ecological models of evolution. Our model describes evolutionary dynamics in a high-dimensional, structured genotype space with a significant role for interspecific interaction. We find some promising qualitative similarity with the empirical facts about macroevolution, including broadly distributed extinction sizes and realistic exploration of the genotype space. The abstraction of our model permits numerous interpretations and applications beyond macroevolution, including molecular evolution and technological innovation.
💡 Research Summary
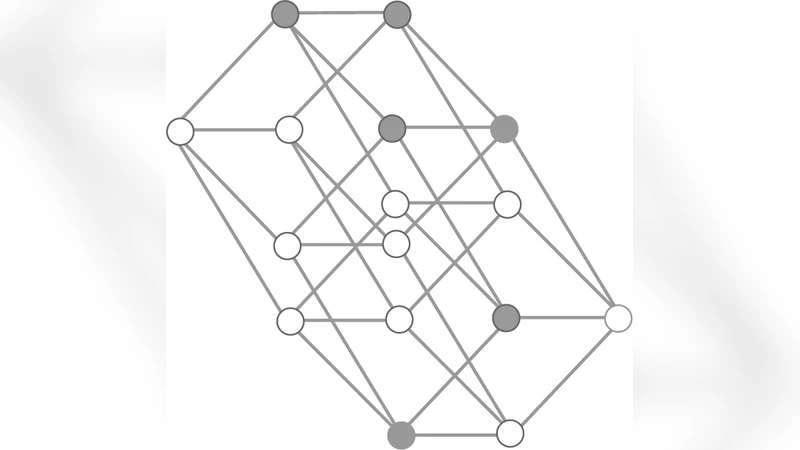
The paper presents a unified, highly abstract model that bridges the classic fitness‑landscape approach and ecological interaction frameworks to explore macro‑evolutionary dynamics. The authors construct a high‑dimensional genotype space represented as a lattice where each node corresponds to a specific genotype. A static fitness value is initially assigned to each node, but unlike traditional fitness‑landscape models, this value is allowed to change dynamically as a function of inter‑specific interactions. These interactions are encoded in a matrix that specifies how the presence of one species at a given node (or a neighboring node) modifies the fitness of another species—competition reduces fitness, mutualism raises it, and neutral relationships leave it unchanged.
Evolution proceeds through two elementary processes: mutation, modeled as a stochastic hop to an adjacent lattice site, and selection, implemented by replicating individuals proportionally to the current fitness of their site. Population size is kept constant by normalizing replication events. The model thus captures a feedback loop: as species occupy certain regions, they reshape the fitness landscape for themselves and others, which in turn drives further movement and replication.
Simulation experiments reveal several emergent phenomena that align with empirical macro‑evolutionary observations. First, the system exhibits punctuated equilibrium: periods of relative stasis are interrupted by abrupt reorganizations of the interaction matrix that cause rapid declines in fitness across many nodes, leading to simultaneous extinctions and the colonization of new genotype regions. Second, extinction event sizes follow a power‑law distribution, indicating many small extinctions and a few very large ones—a hallmark of self‑organized criticality observed in the fossil record. Third, the trajectories through genotype space display fractal‑like properties, mixing gradual, stepwise mutations with occasional large jumps, mirroring the dual nature of molecular evolution (point mutations versus large‑scale rearrangements).
Beyond biological evolution, the authors emphasize the model’s broad applicability. By reinterpreting lattice nodes as technological ideas, cultural memes, or software modules, and redefining the interaction matrix as market competition, social influence, or code dependencies, the same dynamics can simulate waves of innovation, diffusion of cultural traits, and sudden technological collapses. This versatility underscores the value of an abstract, parameter‑light framework for studying complex adaptive systems in general.
In conclusion, the study demonstrates that neither a static fitness landscape nor pure ecological interaction alone can fully account for macro‑evolutionary patterns. Their integration yields a richer dynamical system capable of reproducing realistic extinction statistics, evolutionary bursts, and exploratory behavior in genotype space. The authors suggest future work to calibrate the model against empirical genetic and fossil data, to infer realistic interaction matrices from ecological networks, and to perform sensitivity analyses of key parameters. Such extensions would strengthen the model’s predictive power and further illuminate the intertwined roles of fitness structure and contingency in shaping the history of life and other evolving systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment