Origins of Binary Gene Expression in Post-transcriptional Regulation by MicroRNAs
MicroRNA-mediated regulation of gene expression is characterised by some distinctive features that set it apart from unregulated and transcription factor-regulated gene expression. Recently, a mathematical model has been proposed to describe the dynamics of post-transcriptional regulation by microRNAs. The model explains the observations made in single cell experiments quite well. In this paper, we introduce some additional features into the model and consider two specific cases. In the first case, a non-cooperative positive feedback loop is included in the transcriptional regulation of the target gene expression. In the second case, a stochastic version of the original model is considered in which there are random transitions between the inactive and active expression states of the gene. In the first case we show that bistability is possible in a parameter regime, due to the presence of a non-linear protein decay term in the gene expression dynamics. In the second case, we derive the conditions for obtaining stochastic binary gene expression. We find that this type of gene expression is more favourable in the case of regulation by microRNAs as compared to the case of unregulated gene expression. The theoretical predictions relating to binary gene expression are experimentally testable.
💡 Research Summary
The paper builds on a previously published deterministic model of microRNA‑mediated post‑transcriptional regulation and introduces two major extensions to explain binary (ON/OFF) gene‑expression patterns that have been observed in single‑cell experiments.
First, the authors embed a non‑cooperative positive feedback loop into the transcriptional regulation of the target gene. The transcription rate is taken as α + βP, where P is the protein concentration and β is a linear feedback coefficient. In addition, protein degradation is modeled with a nonlinear term δP + γPⁿ (n > 1), so that degradation accelerates sharply once protein levels exceed a threshold. By performing bifurcation analysis, they show that when β and γ are sufficiently large relative to the basal transcription rate α, the system possesses two stable steady states (low‑ and high‑expression) separated by an unstable saddle point. This bistability arises even though the feedback is non‑cooperative, because the nonlinear decay creates the necessary curvature in the nullclines. The authors also discuss hysteresis: the system’s state depends on its history, a hallmark of switch‑like behavior.
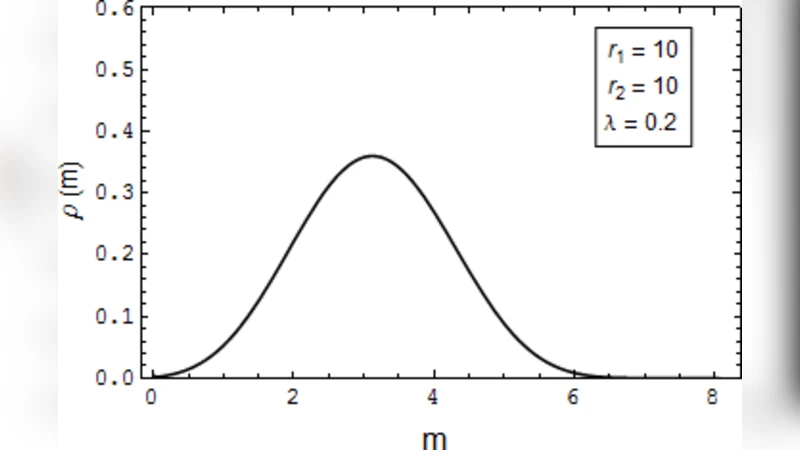
Second, the paper develops a stochastic version of the original scheme. The gene toggles between an inactive (off) state and an active (on) state with transition rates k_off and k_on, respectively. In the on state transcription proceeds at rate r, while in the off state it is zero. The downstream mRNA‑microRNA interaction and translation inhibition are retained from the deterministic model. By writing the master equation for the joint probability distribution of mRNA and protein numbers, the authors derive analytical expressions for the steady‑state distribution. They find that when the ratio k_on/k_off lies within a specific window, the probability distribution becomes bimodal, i.e., two distinct peaks corresponding to low and high protein levels. Crucially, the presence of microRNAs lowers the effective degradation rates of both mRNA and protein, which broadens the window of k_on/k_off values that yield bimodality. Consequently, binary expression is more readily achieved under microRNA regulation than in an unregulated system.
Numerical simulations confirm the analytical predictions. In the deterministic feedback case, simulations display hysteresis loops and switching behavior as parameters are varied. In the stochastic case, increasing microRNA concentration sharpens the two peaks of the protein distribution and raises the probability of observing clear ON/OFF states. Comparisons with a model lacking microRNA regulation show that the latter requires much tighter tuning of k_on and k_off to produce bimodality.
The authors propose experimental tests: (i) fluorescent tagging of both the target mRNA and its regulating microRNA in live single cells, combined with time‑lapse microscopy to monitor transcription bursts and translation events; (ii) CRISPR‑based activation or repression of the target gene to artificially modify the feedback strength β; and (iii) manipulation of microRNA levels using mimics or inhibitors to assess their impact on the bimodal distribution. By measuring the dwell times in ON and OFF states and the shape of the protein‑level histogram, the theoretical conditions derived in the paper can be directly validated.
In summary, the study demonstrates that (1) a non‑cooperative positive feedback combined with nonlinear protein decay can generate bistability in microRNA‑regulated networks, and (2) stochastic switching between transcriptional states is amplified by microRNA‑mediated reduction of degradation, making binary gene expression more favorable. These insights extend current understanding of microRNA function beyond simple repression, highlighting its role in shaping cellular decision‑making and phenotypic heterogeneity.
Comments & Academic Discussion
Loading comments...
Leave a Comment