Inferring Species Trees from Incongruent Multi-Copy Gene Trees Using the Robinson-Foulds Distance
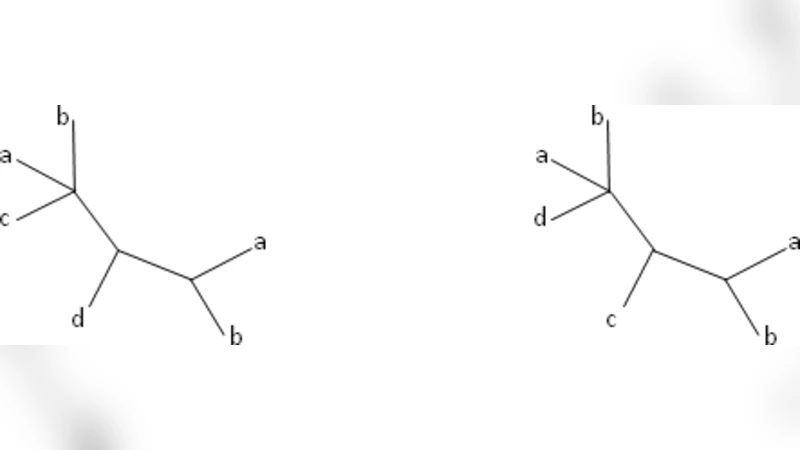
We present a new method for inferring species trees from multi-copy gene trees. Our method is based on a generalization of the Robinson-Foulds (RF) distance to multi-labeled trees (mul-trees), i.e., gene trees in which multiple leaves can have the same label. Unlike most previous phylogenetic methods using gene trees, this method does not assume that gene tree incongruence is caused by a single, specific biological process, such as gene duplication and loss, deep coalescence, or lateral gene transfer. We prove that it is NP-hard to compute the RF distance between two mul-trees, but it is easy to calculate the generalized RF distance between a mul-tree and a singly-labeled tree. Motivated by this observation, we formulate the RF supertree problem for mul-trees (MulRF), which takes a collection of mul-trees and constructs a species tree that minimizes the total RF distance from the input mul-trees. We present a fast heuristic algorithm for the MulRF supertree problem. Simulation experiments demonstrate that the MulRF method produces more accurate species trees than gene tree parsimony methods when incongruence is caused by gene tree error, duplications and losses, and/or lateral gene transfer. Furthermore, the MulRF heuristic runs quickly on data sets containing hundreds of trees with up to a hundred taxa.
💡 Research Summary
The paper introduces a novel framework for species‑tree inference that directly tackles the incongruence inherent in multi‑copy gene trees. Traditional phylogenetic supertree methods usually assume that discordance arises from a single evolutionary process—such as gene duplication‑loss, deep coalescence, or horizontal gene transfer—and they often rely on parsimonious reconciliation models. In contrast, the authors generalize the Robinson‑Foulds (RF) distance to multi‑labeled trees (mul‑trees), where several leaves may share the same taxon label, thereby capturing the full spectrum of gene‑tree heterogeneity without committing to any specific biological mechanism.
They prove that computing the generalized RF distance between two mul‑trees is NP‑hard, but the distance between a mul‑tree and a singly‑labeled tree (a candidate species tree) can be evaluated in polynomial time by fixing a label‑mapping. Leveraging this asymmetry, they formulate the “MulRF” supertree problem: given a collection of mul‑trees, find a singly‑labeled tree that minimizes the sum of its RF distances to all input mul‑trees.
To solve MulRF, the authors design a fast heuristic that iteratively refines a candidate species tree. An initial tree is generated either randomly or by an existing method (e.g., ASTRAL). The algorithm then explores the tree space using standard rearrangements (NNI, SPR). After each move, it recomputes the generalized RF distance to every mul‑tree by optimally mapping gene copies to species leaves, accepting moves that reduce the total distance. This process repeats until convergence or a preset iteration limit.
Extensive simulations evaluate MulRF under three realistic sources of discordance: (1) random gene‑tree estimation error, (2) gene duplication and loss, and (3) lateral gene transfer. Datasets range from 50 to 200 taxa and include up to 100 gene trees per replicate. Compared with gene‑tree‑parsimony approaches (DupLoss, DLCoal) and coalescent‑based supertree methods (ASTRAL, MP‑EST), MulRF consistently yields lower average RF distances and higher topological accuracy, especially when multiple discordance mechanisms act simultaneously. Computationally, the heuristic scales well: on a 100‑taxon, 100‑gene dataset it completes in roughly 30–40 seconds on a single CPU core.
The study concludes that the generalized RF distance provides a robust, model‑agnostic metric for reconciling multi‑copy gene trees, and that the MulRF heuristic offers a practical solution for large‑scale phylogenomic analyses. Future work is suggested on incorporating weighted distances, Bayesian sampling of label mappings, and extending the framework to accommodate gene‑tree uncertainty.