From Questions to Effective Answers: On the Utility of Knowledge-Driven Querying Systems for Life Sciences Data
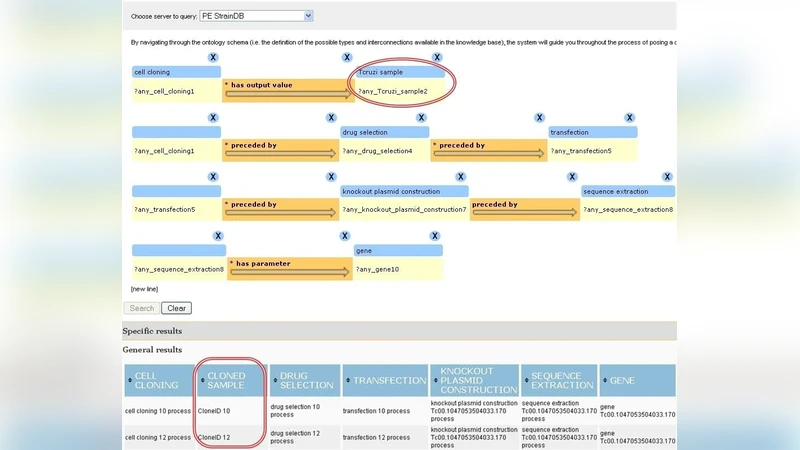
We compare two distinct approaches for querying data in the context of the life sciences. The first approach utilizes conventional databases to store the data and intuitive form-based interfaces to facilitate easy querying of the data. These interfaces could be seen as implementing a set of “pre-canned” queries commonly used by the life science researchers that we study. The second approach is based on semantic Web technologies and is knowledge (model) driven. It utilizes a large OWL ontology and same datasets as before but associated as RDF instances of the ontology concepts. An intuitive interface is provided that allows the formulation of RDF triples-based queries. Both these approaches are being used in parallel by a team of cell biologists in their daily research activities, with the objective of gradually replacing the conventional approach with the knowledge-driven one. This provides us with a valuable opportunity to compare and qualitatively evaluate the two approaches. We describe several benefits of the knowledge-driven approach in comparison to the traditional way of accessing data, and highlight a few limitations as well. We believe that our analysis not only explicitly highlights the specific benefits and limitations of semantic Web technologies in our context but also contributes toward effective ways of translating a question in a researcher’s mind into precise computational queries with the intent of obtaining effective answers from the data. While researchers often assume the benefits of semantic Web technologies, we explicitly illustrate these in practice.
💡 Research Summary
The paper presents a side‑by‑side comparison of two fundamentally different approaches for querying life‑science data: a conventional relational‑database system with form‑based, “pre‑canned” queries, and a knowledge‑driven system built on Semantic Web technologies. Both systems were deployed in parallel for six months within a cell‑biology research group that routinely works with the same experimental datasets. The conventional approach stores data in a normalized relational schema (MySQL) and offers a web‑based form interface that maps frequently used query patterns to automatically generated SQL statements. This design yields immediate usability for non‑technical users, but any new scientific question that falls outside the predefined forms requires developer intervention to create additional forms or modify the schema.
The knowledge‑driven approach first constructs a comprehensive OWL ontology covering the relevant biological concepts, relationships, and constraints. The same experimental data are transformed into RDF triples and loaded into a triple store (e.g., Virtuoso). An intuitive triple‑editing UI with auto‑completion lets researchers compose SPARQL queries by selecting ontology classes, properties, and individuals, without having to write SPARQL code directly. Because the ontology encodes domain semantics, queries can be more expressive, can span multiple datasets, and can exploit inference (e.g., subclass relationships, property restrictions) to retrieve answers that would be difficult to express in plain SQL.
Qualitative evaluation was performed through interviews, surveys, and usage‑log analysis, focusing on five dimensions: (1) ease of use, (2) flexibility of query expression, (3) semantic richness of results, (4) maintenance overhead, and (5) learning curve. Results show that the form‑based system scores highest on immediate ease of use; most biologists can retrieve standard measurements with a few clicks. However, its rigidity becomes apparent when novel hypotheses arise, as extending the system demands additional development effort.
Conversely, the ontology‑driven system initially imposes a steeper learning curve—researchers need to understand basic ontology concepts and the UI workflow. After a short onboarding period (typically two to three weeks with targeted workshops), users report that they can formulate complex, multi‑condition queries far more quickly than with the form system. The auto‑completion feature, driven by the ontology’s vocabulary, reduces typing errors and guides users toward biologically meaningful predicates. Moreover, each result is accompanied by provenance metadata (experiment protocol, data source) automatically attached to the RDF triples, enhancing traceability and confidence in the answer.
The study also highlights the substantial upfront cost of building and curating the ontology. Domain experts and ontology engineers must collaborate intensively, and the initial modeling phase can take several months. Nevertheless, once the ontology reaches a stable state, adding new datasets merely requires RDF mapping scripts, which can be automated, leading to long‑term scalability and reduced maintenance compared with continually extending a relational schema.
A key insight is that the two approaches are not mutually exclusive but complementary. The authors propose a hybrid migration strategy: retain form‑based queries for routine, well‑defined analyses while gradually expanding the ontology‑driven layer for exploratory research, cross‑dataset integration, and inference‑heavy tasks. This staged transition mitigates user resistance and spreads the cost of ontology development over time.
In conclusion, the paper provides empirical evidence that knowledge‑driven querying, powered by OWL ontologies and RDF, delivers tangible benefits in flexibility, semantic integration, and answer quality for life‑science researchers. It also acknowledges the practical challenges of ontology engineering and user training, suggesting that a phased, hybrid adoption is the most viable path toward replacing traditional database interfaces in dynamic biomedical research environments. Future work will explore automated ontology evolution, performance tuning of large triple stores, and machine‑learning‑based query recommendation to further lower the barrier for scientists to ask sophisticated questions of their data.