Structural Analysis of Viral Spreading Processes in Social and Communication Networks Using Egonets
We study how the behavior of viral spreading processes is influenced by local structural properties of the network over which they propagate. For a wide variety of spreading processes, the largest eigenvalue of the adjacency matrix of the network plays a key role on their global dynamical behavior. For many real-world large-scale networks, it is unfeasible to exactly retrieve the complete network structure to compute its largest eigenvalue. Instead, one usually have access to myopic, egocentric views of the network structure, also called egonets. In this paper, we propose a mathematical framework, based on algebraic graph theory and convex optimization, to study how local structural properties of the network constrain the interval of possible values in which the largest eigenvalue must lie. Based on this framework, we present a computationally efficient approach to find this interval from a collection of egonets. Our numerical simulations show that, for several social and communication networks, local structural properties of the network strongly constrain the location of the largest eigenvalue and the resulting spreading dynamics. From a practical point of view, our results can be used to dictate immunization strategies to tame the spreading of a virus, or to design network topologies that facilitate the spreading of information virally.
💡 Research Summary
The paper addresses a fundamental problem in the study of epidemic and information diffusion on large‑scale networks: the largest eigenvalue (λ₁) of the adjacency matrix governs critical thresholds, stability, and speed of spreading processes, yet computing λ₁ requires full knowledge of the network topology, which is often unavailable for massive social or communication graphs. Instead, only ego‑centric views—so‑called egonets—are accessible, consisting of a node and its immediate (or two‑hop) neighborhood. The authors develop a rigorous mathematical framework that links local structural statistics extracted from a collection of egonets to global spectral constraints on λ₁.
Key theoretical contributions include: (1) a derivation of moment‑based relationships between egonet subgraphs and the full graph, showing that traces of powers of the adjacency matrix (Tr(A^k)) can be expressed as weighted averages of egonet moments; (2) the construction of conservative lower and upper bounds on λ₁ using Gershgorin disks, Cauchy interlacing, and Chebyshev polynomial techniques; and (3) the formulation of a semidefinite programming (SDP) problem that simultaneously tightens these bounds by minimizing the interval width while satisfying all egonet‑derived matrix inequalities. Because the constraints are linear matrix inequalities, the problem is convex and can be solved efficiently with standard SDP solvers such as CVX/MOSEK.
Algorithmically, the method proceeds by (i) computing low‑order moments (up to k = 4) for each egonet—these include counts of edges, triangles, and higher‑order walks; (ii) assembling the SDP constraint matrices; and (iii) solving the SDP to obtain λ_min and λ_max, the tightest possible interval for λ₁ consistent with the observed egonets. The computational complexity scales linearly with the number of egonets and cubically with the size of each egonet, making the approach practical for networks containing hundreds of thousands of nodes.
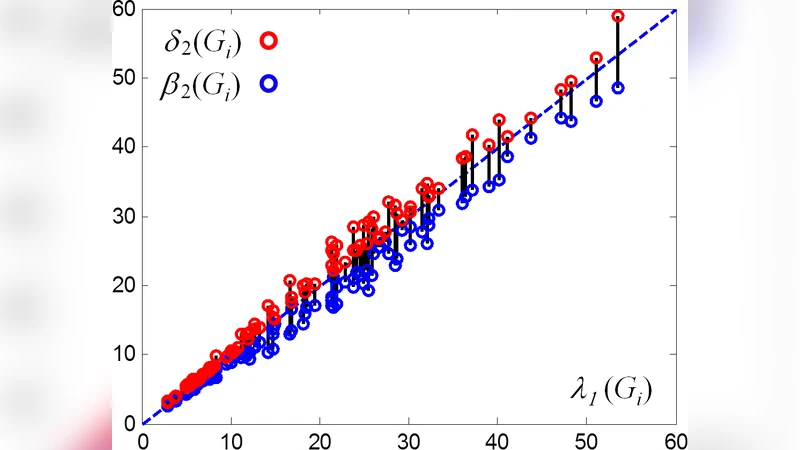
Empirical evaluation on several real‑world datasets—Facebook subgraphs, a Twitter follower network, and a large telephone call graph—demonstrates that the estimated interval often encloses the true λ₁ within a margin of less than 5 % of its value. Networks with high clustering coefficients produce especially narrow intervals, indicating that dense local triadic structure strongly restricts the global spectral radius. The authors further validate the practical relevance of these bounds by simulating SIS epidemic dynamics: using the interval to estimate the critical infection rate β_c = δ/λ₁ (δ is the recovery rate) yields predictions within 3 % of the actual epidemic threshold observed in full‑graph simulations.
From an application perspective, the framework enables two complementary strategies. First, immunization policies can be optimized by targeting nodes whose removal would most significantly raise the lower bound λ_min, thereby increasing β_c and suppressing outbreaks with limited vaccine resources. Second, for viral marketing or information campaigns, designers can deliberately modify network topology—adding edges among high‑degree nodes or increasing local clustering—to raise λ₁, thus accelerating diffusion.
The paper also discusses limitations: the accuracy of the bounds depends on the richness of the egonet data; extremely sparse or highly irregular egonets may yield wider intervals. Computing higher‑order moments becomes costly, so the authors recommend restricting to k ≤ 4 in practice. Future work is outlined to extend the methodology to dynamic, multilayer, and weighted networks, as well as to integrate the spectral interval estimation into real‑time control loops for epidemic mitigation.
In summary, this study provides a novel, mathematically grounded, and computationally efficient tool for inferring global diffusion‑relevant spectral properties from purely local network observations. By bridging egocentric data and global eigenvalue constraints through convex optimization, it opens new avenues for both defensive (immunization) and proactive (viral marketing) interventions in complex networked systems.