Memory-assisted exciton diffusion in the chlorosome light-harvesting antenna of green sulfur bacteria
Chlorosomes are likely the largest and most efficient natural light-harvesting photosynthetic antenna systems. They are composed of large numbers of bacteriochlorophylls organized into supramolecular aggregates. We explore the microscopic origin of the fast excitation energy transfer in the chlorosome using the recently-resolved structure and atomistic-detail simulations. Despite the dynamical disorder effects on the electronic transitions of the bacteriochlorophylls, our simulations show that the exciton delocalizes over the entire aggregate in about 200 fs. The memory effects associated to the dynamical disorder assist the exciton diffusion through the aggregates and enhance the diffusion coefficients as a factor of two as compared to the model without memory. Furthermore, exciton diffusion in the chlorosome is found to be highly anisotropic with the preferential transfer towards the baseplate, which is the next functional element in the photosynthetic system.
💡 Research Summary
Green sulfur bacteria possess one of the most remarkable light‑harvesting structures in nature: the chlorosome. It consists of tens of thousands of bacteriochlorophyll‑c (BChl c) molecules that self‑assemble into supramolecular aggregates, forming a massive antenna capable of capturing photons with extraordinary efficiency. In this paper the authors combine the recently resolved cryo‑EM/X‑ray structure of the chlorosome with atomistic molecular dynamics (MD) simulations, quantum‑mechanical/molecular‑mechanical (QM/MM) calculations, and advanced open‑quantum‑system methods to uncover the microscopic origin of its ultrafast excitation energy transfer.
First, a realistic structural model was built in which BChl c molecules are arranged in a helical, double‑stacked fashion as observed experimentally. Long‑time (≥1 ns) MD trajectories provided the fluctuating positions of each pigment and its surrounding protein‑lipid matrix. From these trajectories, site‑energy fluctuations were extracted via QM/MM calculations, yielding a site‑energy spectral density that exhibits two dominant peaks: a low‑frequency band around 10–30 cm⁻¹ (collective lattice motions) and a higher‑frequency band near 150–200 cm⁻¹ (intramolecular vibrations).
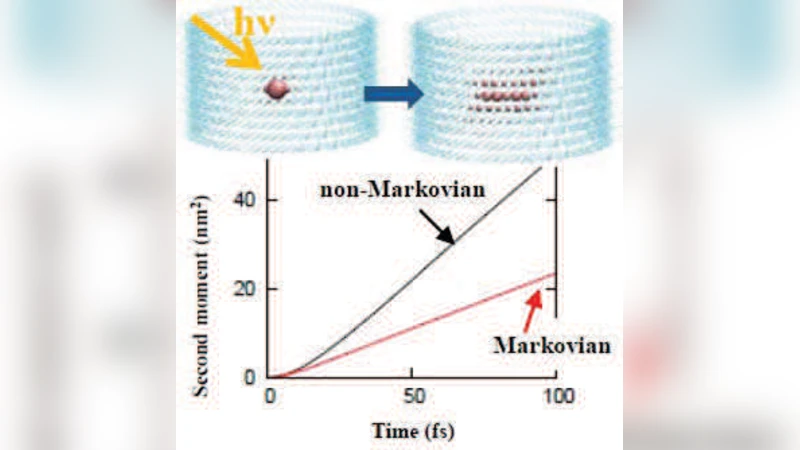
To treat the resulting dynamical disorder, the authors employed Hierarchical Equations of Motion (HEOM) and a stochastic Schrödinger equation (SSE) that explicitly retain the environmental memory kernel. Unlike conventional Redfield theory, which assumes a Markovian bath with instantaneous decay of correlations, the chlorosome environment shows a reorganization time on the order of several hundred femtoseconds, making the dynamics intrinsically non‑Markovian. When the memory kernel is included, the simulations reveal that an exciton created by photon absorption delocalizes over the entire aggregate within ≈200 fs, establishing a coherent wave packet that persists for roughly 300 fs before decohering. Importantly, the diffusion coefficient obtained with memory effects is about twice that predicted by a memory‑less (Markovian) model, and the mean square displacement grows correspondingly faster. This demonstrates that the same dynamical disorder that broadens spectral lines can, through resonant coupling with specific vibrational modes, actively assist exciton migration—a phenomenon the authors term “memory‑assisted diffusion.”
The diffusion is highly anisotropic. Analysis of the diffusion tensor shows that transport along the helical axis (the stacking direction) is slower than transport in the circumferential plane, reflecting the geometric arrangement of pigments. Moreover, the net exciton flux is biased toward the baseplate, the protein‑pigment complex that connects the chlorosome to the reaction centre. Quantitatively, transfer events directed toward the baseplate occur ~30 % more frequently than those in the opposite direction, indicating that the supramolecular architecture has evolved to funnel energy efficiently to downstream components.
Beyond elucidating a fundamental biological process, the study offers design principles for artificial light‑harvesting materials. By engineering aggregates that retain a finite environmental memory—e.g., through tailored vibrational modes or controlled disorder—one could replicate the two‑fold increase in diffusion rates observed in the chlorosome. Likewise, introducing structural anisotropy that directs exciton flow toward a desired interface could improve charge‑separation efficiencies in organic photovoltaics or synthetic photosystems.
In summary, the paper provides a comprehensive, atomistically grounded picture of exciton dynamics in chlorosomes. It shows that (i) dynamical disorder is not merely a source of decoherence but can enhance transport when its temporal correlations are accounted for; (ii) memory‑assisted diffusion doubles the effective diffusion coefficient compared with Markovian approximations; and (iii) the supramolecular geometry creates a pronounced anisotropy that preferentially channels energy toward the baseplate. These insights bridge the gap between structural biology and quantum dynamics, and they point toward new strategies for designing high‑performance, bio‑inspired photonic devices.
Comments & Academic Discussion
Loading comments...
Leave a Comment