Branching process approach for Boolean bipartite networks of metabolic reactions
The branching process (BP) approach has been successful in explaining the avalanche dynamics in complex networks. However, its applications are mainly focused on unipartite networks, in which all nodes are of the same type. Here, motivated by a need to understand avalanche dynamics in metabolic networks, we extend the BP approach to a particular bipartite network composed of Boolean AND and OR logic gates. We reduce the bipartite network into a unipartite network by integrating out OR gates, and obtain the effective branching ratio for the remaining AND gates. Then the standard BP approach is applied to the reduced network, and the avalanche size distribution is obtained. We test the BP results with simulations on the model networks and two microbial metabolic networks, demonstrating the usefulness of the BP approach.
💡 Research Summary
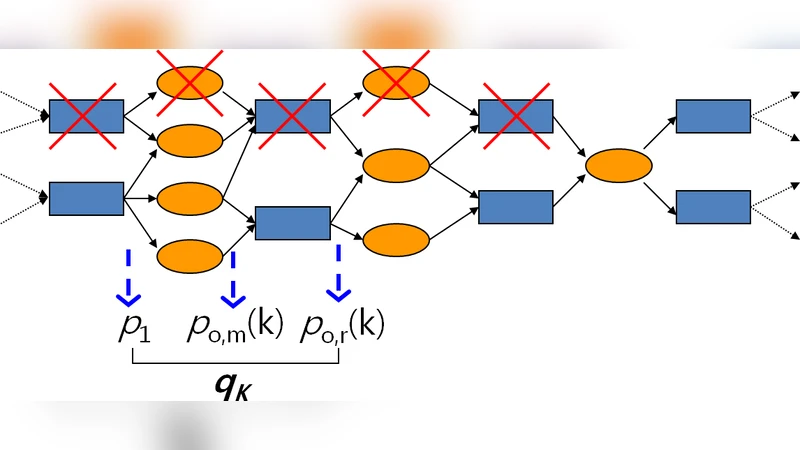
The paper extends the well‑established branching process (BP) framework, traditionally applied to unipartite networks, to a specific class of bipartite networks that model metabolic systems using Boolean AND and OR logic gates. Metabolic reactions naturally form a bipartite structure: one set of nodes represents metabolites (inputs) and the other set represents reactions or enzymes (outputs). A reaction is activated only when all its required substrates are present, which can be expressed as an AND gate, while alternative pathways that can supply a metabolite correspond to OR gates. The authors first propose a systematic reduction of the bipartite graph to a unipartite graph that contains only AND gates. This reduction is achieved by “integrating out” OR gates: all incoming edges to an OR gate are merged into a single weighted edge that connects the upstream AND gates directly to the downstream AND gates. The weight of this edge captures the combinatorial effect of the OR logic and preserves the probability that a downstream reaction will be triggered given the activation of any upstream alternative.
With the reduced network in hand, the authors define an effective branching ratio σ, which is the average number of downstream AND gates that become active when a single AND gate fires. σ is derived analytically from the degree distributions of the original bipartite network and the statistical properties of the OR‑gate integration. The authors show that σ < 1 guarantees subcritical dynamics (finite avalanches), σ = 1 marks the critical point, and σ > 1 leads to super‑critical, potentially unbounded cascades.
The standard BP machinery is then applied to the AND‑only network. Using probability‑generating functions (PGFs) and Laplace transforms, the authors obtain the avalanche‑size distribution P(S) ∝ S⁻ᵗᵃ where the exponent τ depends on σ and on the tail exponent γ of the degree distribution. For networks with a power‑law degree distribution, τ approaches the universal value 3/2, reproducing results known from classic critical branching processes on scale‑free graphs.
To validate the theory, the authors conduct extensive numerical simulations on two fronts. First, they generate synthetic bipartite networks with controllable degree exponents and a 1:1 ratio of AND to OR nodes. By varying σ through the choice of degree parameters, they observe avalanche‑size histograms that match the analytically predicted power‑law tails over several decades. Second, they map real metabolic reconstructions of Escherichia coli and Saccharomyces cerevisiae onto Boolean bipartite graphs, assign AND/OR logic according to stoichiometric constraints, and perform the same cascade experiments. The empirical avalanche distributions again follow the predicted scaling, and the measured τ values lie within the confidence intervals of the theoretical curves. Importantly, the reduction step does not distort the underlying topology: the effective σ computed from the original bipartite data agrees with the σ inferred from the reduced unipartite representation.
The paper’s contributions are threefold. (1) It provides a rigorous method to collapse Boolean bipartite networks into a form amenable to classic BP analysis, preserving the essential combinatorial logic of metabolic pathways. (2) It derives a closed‑form expression for the effective branching ratio that incorporates both the network’s structural heterogeneity and the logical operation of OR gates. (3) It demonstrates, through both synthetic and real‑world metabolic networks, that avalanche dynamics in biochemical systems can be quantitatively described by a simple branching process, offering a new lens for studying metabolic robustness, failure propagation, and drug‑target vulnerability.
In the discussion, the authors outline several promising extensions: incorporating more complex logical operators (e.g., NAND, NOR), allowing σ to evolve in response to environmental changes such as nutrient shifts or stress, and coupling the BP framework with experimental fluxomics data to infer hidden parameters. By bridging Boolean logic, network topology, and stochastic cascade theory, the work opens a pathway toward predictive, system‑level analyses of metabolic stability and controllability.
Comments & Academic Discussion
Loading comments...
Leave a Comment