A New Approach to Understanding Ontogenesis and The Theory of Aging
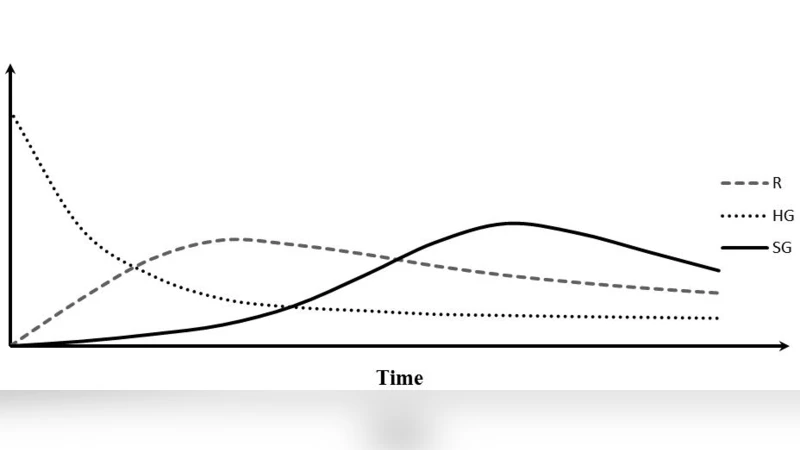
This paper proposes an original theory of aging of multicellular organisms. The cells of multicellular organisms, in contrast to unicellular organisms, are burdened with a two- part genome: housekeeping and specialized (multicellular), responsible for ontogenesis and terminal differentiation. The two parts of the genome compete for limited adaptive resources thereby interfering with the ability of the house-keeping part of the genome to adequately perform reparative and adaptive functions in post mitotic cells. The necessity to complete the ontgenesis program, leads to increased activity of the multicellular components of the genome. As a result, the allocation of cellular resources to specialized genome con-tinuously increases with time. This leads to a deficit of reparative and adaptive capacity in post mitotic cells. Suggestions for future research focus on identifying groups of genes responsible for regulation of growth rate of specialized genome and suppressing ability of the cell division. A better understanding of the relationship between the two parts of the genome will not only help us to manipulate ontogenesis and aging, but will also improve our understanding of cancer development and ontogenesis.
💡 Research Summary
The paper puts forward a novel conceptual framework for aging in multicellular organisms that hinges on the idea of a “dual‑part” genome. The authors divide the cellular genome into a housekeeping component, responsible for core metabolic processes, DNA repair, protein quality control, and stress responses, and a specialized, multicellular component that drives ontogeny, tissue differentiation, growth signaling, and, indirectly, tumor suppression. During early development the specialized genome is massively up‑regulated to orchestrate organogenesis; this surge consumes a substantial share of limited intracellular resources such as ATP, amino acids, nucleotides, and enzymatic capacity. Because the total pool of these resources is finite, the increasing demand of the specialized genome inevitably reduces the allocation available to the housekeeping genome.
The authors argue that this competition becomes especially detrimental in post‑mitotic cells (neurons, myocytes, cardiomyocytes), which rely heavily on housekeeping functions for long‑term maintenance. As the specialized genome continues to draw resources, the efficiency of DNA repair enzymes, antioxidant systems, and proteostasis networks declines. Accumulated DNA lesions, mitochondrial dysfunction, protein aggregation, and chronic low‑grade inflammation then emerge, manifesting as the physiological hallmarks of aging. In this view, aging is not merely the passive accrual of damage but an active, resource‑driven shift in genomic priority that favors developmental programs at the expense of cellular maintenance.
The paper also links this mechanism to cancer biology. Tumor cells often reactivate developmental pathways and bypass cell‑cycle checkpoints, effectively hijacking the specialized genome’s growth‑promoting machinery while simultaneously suppressing housekeeping safeguards. Thus, the same genomic competition that drives aging may also predispose tissues to malignant transformation when regulatory balances break down.
Strengths of the work include its integrative perspective—bringing together developmental biology, metabolic economics, and aging theory—and its bold hypothesis that a single underlying resource competition could explain multiple age‑related phenomena. By framing ontogeny and senescence as two ends of a continuum, the authors open a conceptual space for interventions that could simultaneously extend healthspan and reduce cancer risk.
However, the manuscript falls short on empirical validation. No quantitative data are presented to demonstrate actual resource reallocation between the two genomic sectors, nor are specific gene sets identified that constitute the “growth‑rate regulators” of the specialized genome. The model assumes a uniform resource pool across diverse cell types, ignoring tissue‑specific metabolic specializations and differential reliance on housekeeping functions. Moreover, the proposal to suppress cell‑division ability as an anti‑aging strategy conflicts with the essential role of controlled proliferation in tissue repair and immune function.
Future research directions suggested by the authors are appropriate but need refinement. High‑resolution single‑cell RNA‑seq combined with metabolomic flux analysis could map the dynamic balance of housekeeping versus specialized transcriptomes and their associated energy consumption over the lifespan. CRISPR‑based perturbation of candidate regulators (e.g., master transcription factors, epigenetic modifiers) would allow testing whether dampening specialized‑genome activity restores repair capacity and delays functional decline in post‑mitotic tissues. Parallel studies in oncogenic models could assess whether the same interventions reduce tumor initiation by re‑establishing housekeeping dominance.
If these experimental avenues bear fruit, the dual‑genome competition model could become a unifying principle linking development, aging, and cancer, offering novel therapeutic targets that modulate the internal allocation of cellular resources rather than merely addressing downstream damage.
Comments & Academic Discussion
Loading comments...
Leave a Comment