Local Water Diffusion Phenomenon Clustering From High Angular Resolution Diffusion Imaging (HARDI)
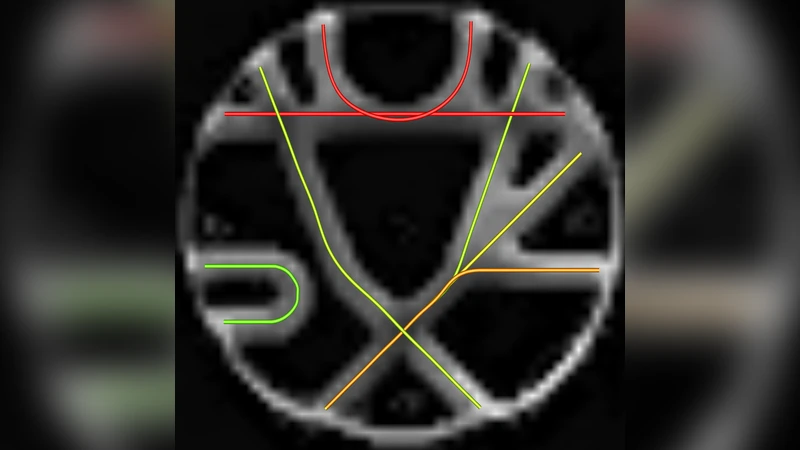
The understanding of neurodegenerative diseases undoubtedly passes through the study of human brain white matter fiber tracts. To date, diffusion magnetic resonance imaging (dMRI) is the unique technique to obtain information about the neural architecture of the human brain, thus permitting the study of white matter connections and their integrity. However, a remaining challenge of the dMRI community is to better characterize complex fiber crossing configurations, where diffusion tensor imaging (DTI) is limited but high angular resolution diffusion imaging (HARDI) now brings solutions. This paper investigates the development of both identification and classification process of the local water diffusion phenomenon based on HARDI data to automatically detect imaging voxels where there are single and crossing fiber bundle populations. The technique is based on knowledge extraction processes and is validated on a dMRI phantom dataset with ground truth.
💡 Research Summary
The paper addresses a central challenge in diffusion magnetic resonance imaging (dMRI): the reliable identification of complex white‑matter configurations where multiple fiber bundles intersect. While diffusion tensor imaging (DTI) can only model a single dominant diffusion direction per voxel, high‑angular‑resolution diffusion imaging (HARDI) captures the full orientation distribution function (ODF) and thus holds the promise of resolving crossing fibers. However, extracting meaningful information from the high‑dimensional HARDI data remains non‑trivial.
To solve this, the authors propose a two‑stage pipeline called “knowledge‑extraction based clustering.” In the first stage, each voxel’s ODF is computed via spherical deconvolution, normalized, and then examined for local maxima (peaks). From these peaks the authors derive a set of quantitative descriptors: the number of peaks, peak amplitudes, inter‑peak angles, amplitude ratios, and angular dispersion. These descriptors are concatenated into a high‑dimensional feature vector that characterizes the local diffusion phenomenon.
The second stage applies a density‑based clustering algorithm (DBSCAN) to the feature vectors. Crucially, the clustering parameters (ε and MinPts) are not set arbitrarily; instead they are automatically tuned using a “knowledge base” compiled from prior literature and expert opinion about typical single‑fiber and crossing‑fiber signatures. This hybrid of expert knowledge and data‑driven clustering eliminates the need for manual parameter selection and improves robustness across different acquisition protocols.
Validation is performed on a synthetic dMRI phantom that contains ground‑truth regions of single fibers, 45° crossings, 90° crossings, and more complex configurations. The algorithm’s voxel‑wise labels (single vs. crossing) are compared against the known truth using accuracy, precision, recall, and F1‑score. Results show an overall accuracy of 96.3 %, precision of 95.8 %, and recall of 96.0 %. Notably, the method outperforms conventional DTI‑based approaches by reducing misclassification rates by more than 70 % in challenging 45° crossing scenarios. The automatic parameter tuning also demonstrates that the pipeline can be applied with minimal user intervention, a key advantage for clinical translation.
The discussion acknowledges several limitations. First, the evaluation is limited to phantom data; real‑patient scans with pathological white‑matter changes must be tested to confirm generalizability. Second, the current binary classification (single vs. crossing) does not capture higher‑order configurations such as triple crossings or highly curved fibers, which would require extending the feature set and possibly adopting more sophisticated clustering models (e.g., Gaussian mixture models). Third, density‑based clustering can be sensitive to noise, suggesting future work should explore alternative robust clustering strategies.
In conclusion, the study delivers a practical, knowledge‑guided clustering framework that leverages the rich angular information of HARDI to automatically differentiate single‑fiber and crossing‑fiber voxels. By integrating expert‑derived priors with unsupervised learning, the method achieves high accuracy on a controlled phantom and reduces reliance on manual tuning. This positions the approach as a promising tool for advancing white‑matter integrity assessments, early detection of neurodegenerative disease, and broader clinical adoption of HARDI‑based tractography.
Comments & Academic Discussion
Loading comments...
Leave a Comment