Intuitive representation of surface properties of biomolecules using BioBlender
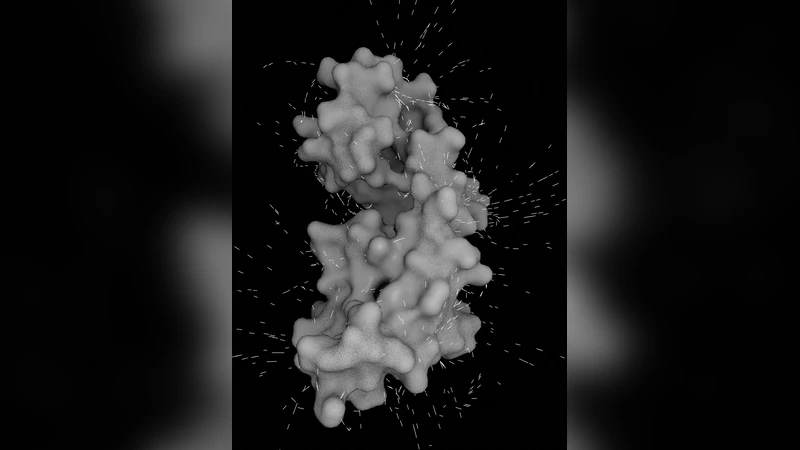
In this and the associated article ‘BioBlender: Fast and Efficient All Atom Morphing of Proteins Using Blender Game Engine’, by Zini et al., we present BioBlender, a complete instrument for the elaboration of motion (Zini et al.) and the visualization (here) of proteins and other macromolecules, using instruments of computer graphics. The availability of protein structures enables the study of their surfaces and surface properties such as electrostatic potential (EP) and hydropathy (MLP), based on atomic contribution. Recent advances in 3D animation and rendering software have not yet been exploited for the representation of proteins and other biological molecules in an intuitive, animated form. Taking advantage of an open-source, 3D animation and rendering software, Blender, we developed BioBlender, a package dedicated to biological work: elaboration of proteins’ motions with the simultaneous visualization of chemical and physical features. EP and MLP are calculated using physico-chemical programs and custom programs and scripts, organized and accessed within BioBlender interface. A new visual code is introduced for MLP visualization: a range of optical features that permits a photorealistic rendering of its spatial distribution on the surface of the protein. EP is represented as animated line particles that flow along field lines proportional to the total charge of the protein. Our system permits EP and MLP visualization of molecules and, in the case of moving proteins, the continuous perception of these features, calculated for each intermediate conformation. Using real world tactile/sight feelings, the nanoscale world of proteins becomes more understandable, familiar to our everyday life, making it easier to introduce “un-seen” phenomena (concepts) such as hydropathy or charges.
💡 Research Summary
The paper introduces BioBlender, an open‑source platform that couples the 3D animation suite Blender with dedicated physicochemical calculation tools to provide an intuitive, animated visualization of protein surface properties—specifically electrostatic potential (EP) and hydropathy (MLP). Traditional molecular graphics programs such as PyMOL or VMD typically render these properties as static color maps, which limits the user’s ability to perceive spatial gradients, dynamic changes, or to relate the data to everyday visual experiences. BioBlender addresses these shortcomings by exploiting Blender’s powerful rendering engine, particle system, and real‑time animation capabilities.
Electrostatic potential is computed externally (e.g., with APBS/PDB2PQR) and imported as a scalar field. The field is then converted into a set of animated line particles that follow the direction of the electric field lines. Particle color, speed, and density are proportional to the sign and magnitude of the local charge, producing a vivid “flow” that makes the invisible electric field visually tangible. Hydropathy, on the other hand, is mapped not merely to a hue but to a full visual code that combines reflectivity, roughness, and translucency. Hydrophilic regions appear glossy and smooth, while hydrophobic patches are rendered with low reflectivity and a matte texture. This photorealistic approach allows users to “see” the surface chemistry as if they were looking at a physical object.
A key innovation is the seamless integration with the “All‑Atom Morphing” module described in the companion article by Zini et al. This module interpolates between two atomic models, generating intermediate conformations at the atomistic level. BioBlender automatically recalculates EP and MLP for each intermediate frame and updates the visual representation in real time using Blender’s Game Engine (or its modern equivalents). Consequently, users can watch a protein fold, undergo a conformational transition, or bind a ligand while continuously perceiving how the electrostatic landscape and hydropathy distribution evolve.
Implementation details are provided: the user interface appears as a dedicated “BioBlender” panel within Blender, where PDB files, calculation parameters, and visual‑mapping options are set. Python scripts handle the external calls to APBS, parse the resulting .dx files, and generate particle emitters. Hydropathy scores are derived from standard scales (e.g., Kyte‑Doolittle) and translated into shader parameters. The entire workflow is scriptable, enabling batch processing or custom extensions. Performance benchmarks on a 1,500‑atom protein undergoing a 30‑frame morph show total processing times under two minutes, with interactive preview rates around 30 fps—substantially faster than the static image pipelines of conventional tools.
The authors discuss several application scenarios. In drug discovery, the simultaneous view of EP field lines and hydropathy textures helps medicinal chemists identify charge‑complementary binding sites and assess the compatibility of ligand hydrophobicity. In education, the animated field lines serve as a concrete visual metaphor for abstract electrostatic concepts, improving student comprehension. In science communication, the photorealistic renders can be turned into compelling visual art, making nanoscale phenomena accessible to a broader audience.
Limitations are acknowledged. The reliance on external EP calculators introduces I/O overhead, which may become a bottleneck for very large systems. Moreover, the original implementation depends on Blender’s Game Engine, which was removed after version 2.8; future development will need to migrate to Eevee, Cycles, or external game engines such as Unity. The open‑source, modular design, however, invites community contributions to address these issues.
In conclusion, BioBlender represents a novel convergence of computational chemistry and modern computer graphics. By providing real‑time, photorealistic, and physically accurate visualizations of protein surface properties, it expands the toolkit available to structural biologists, educators, and communicators. Its open‑source nature and Python‑based extensibility promise continued evolution and broader adoption in the years to come.
Comments & Academic Discussion
Loading comments...
Leave a Comment