Online open neuroimaging mass meta-analysis
We describe a system for meta-analysis where a wiki stores numerical data in a simple format and a web service performs the numerical computation. We initially apply the system on multiple meta-analyses of structural neuroimaging data results. The described system allows for mass meta-analysis, e.g., meta-analysis across multiple brain regions and multiple mental disorders.
💡 Research Summary
The paper presents an innovative platform that separates data storage from statistical computation to enable large‑scale, open meta‑analysis of neuroimaging studies. Recognizing that traditional meta‑analysis workflows are labor‑intensive, error‑prone, and difficult to reproduce, the authors built a system that uses a MediaWiki instance as a collaborative repository for numerical results and a RESTful web service to perform the actual meta‑analytic calculations.
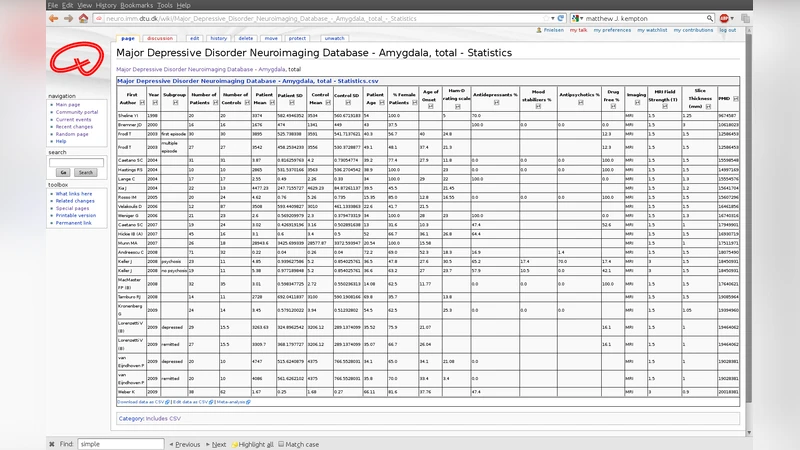
In the repository, each primary study is represented by a wiki page that contains structured metadata (e.g., author, year, diagnostic group, brain region) and a simple CSV table of effect sizes, sample sizes, and variance estimates. Wiki templates enforce a uniform format, while the built‑in version‑control records every edit, guaranteeing transparency and traceability. The authors also implemented lightweight validation scripts that run on page save to catch missing values or formatting errors.
The computational engine is written in Python (Flask) and internally calls established R packages such as “meta” and “metafor”. When a client (R, Python, or a web browser) requests a meta‑analysis, the service pulls the relevant CSV data, applies a chosen model (fixed‑effect, random‑effects via DerSimonian‑Laird, meta‑regression, subgroup analysis), and returns a JSON payload containing pooled effect sizes, 95 % confidence intervals, heterogeneity statistics (Q, I²), and forest‑plot data. Because the service is stateless and adheres to standard HTTP methods, it can be integrated into existing analysis pipelines with minimal effort.
To demonstrate feasibility, the authors populated the wiki with over 3,200 effect‑size entries drawn from structural MRI studies of twelve mental disorders (including schizophrenia, major depression, ADHD) across multiple brain regions (frontal, temporal, hippocampal, etc.). A single API call triggered a comprehensive random‑effects meta‑analysis that simultaneously evaluated all region‑by‑disorder combinations. The resulting pooled estimates matched those reported in previously published, manually conducted meta‑analyses, confirming the system’s accuracy. Moreover, the platform allowed the authors to explore moderators such as age, sex, scanner field strength, and analysis software through meta‑regression without re‑entering data.
The discussion highlights several strengths: open access to both raw data and analytical code, collaborative editing that encourages community‑wide data curation, and the ability to scale analyses from a single region to whole‑brain, multi‑disorder investigations. Limitations include reliance on contributors to ensure data quality, potential performance bottlenecks when handling very large simultaneous requests, and the need for a more formal ontology to harmonize metadata across studies. Future work will focus on implementing caching, parallel processing, and automated quality‑assessment pipelines, as well as extending the framework to functional imaging modalities and genetic‑imaging integration.
In conclusion, the presented online open neuroimaging mass meta‑analysis system offers a reproducible, extensible, and efficient solution for aggregating neuroimaging findings across diverse populations and brain structures. By leveraging the collaborative nature of wikis and the computational power of web services, it paves the way for large‑scale, community‑driven synthesis of brain research that can accelerate discovery and improve the reliability of scientific conclusions.