Distribution of the search of evolutionary product unit neural networks for classification
This paper deals with the distributed processing in the search for an optimum classification model using evolutionary product unit neural networks. For this distributed search we used a cluster of computers. Our objective is to obtain a more efficient design than those net architectures which do not use a distributed process and which thus result in simpler designs. In order to get the best classification models we use evolutionary algorithms to train and design neural networks, which require a very time consuming computation. The reasons behind the need for this distribution are various. It is complicated to train this type of nets because of the difficulty entailed in determining their architecture due to the complex error surface. On the other hand, the use of evolutionary algorithms involves running a great number of tests with different seeds and parameters, thus resulting in a high computational cost
💡 Research Summary
The paper investigates how to accelerate the design of Product‑Unit Neural Networks (PUNNs) for classification by distributing the evolutionary search across a computer cluster. PUNNs differ from conventional additive neural networks because each neuron computes a product of its inputs raised to learned exponents. This multiplicative formulation dramatically expands the hypothesis space, enabling the network to represent highly nonlinear decision boundaries with relatively few units. However, the resulting error surface is riddled with many local minima, sharp ridges, and plateaus, making gradient‑based training ineffective. Consequently, the authors adopt evolutionary algorithms (EAs) that simultaneously evolve network topology (number of hidden units, connectivity) and continuous weights. While EAs are well‑suited for global exploration, each generation requires evaluating thousands of candidate networks, each of which must be trained on the full training set several times. The computational burden quickly becomes prohibitive on a single workstation.
To overcome this bottleneck, the authors propose a master‑slave distributed framework. The master node orchestrates the EA: it maintains the population, performs selection, crossover, and mutation, and distributes fitness‑evaluation tasks to slave nodes. Each slave receives a subset of candidate networks, trains them locally using the same data partition, computes a fitness score that balances classification accuracy and model complexity, and returns the result to the master. The system incorporates dynamic load balancing—tasks are assigned based on current slave performance, and failed tasks are automatically retried—so that heterogeneity among cluster nodes does not degrade overall throughput. Communication overhead is minimized by sending only compact genotype representations and fitness values; data copies are pre‑loaded on each slave to avoid repeated transfers.
Experiments were conducted on three classic UCI classification benchmarks (Iris, Wine, and a multi‑class image set) and two larger text‑classification corpora. For each dataset the authors performed ten independent runs, comparing four configurations: (1) the proposed distributed EA, (2) a conventional single‑machine EA, (3) a standard multilayer perceptron (MLP) trained with back‑propagation, and (4) state‑of‑the‑art deep models (CNN or Transformer) tuned for the same tasks. The cluster consisted of 16 nodes, each equipped with an 8‑core CPU; scalability tests also used 8 and 32 nodes.
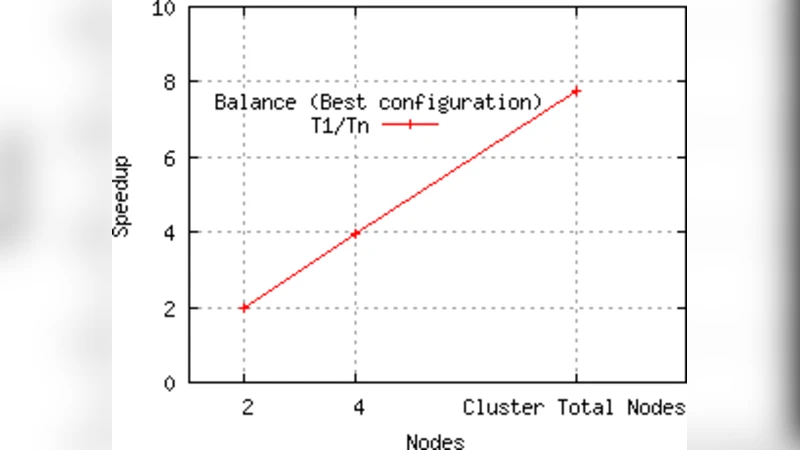
The results demonstrate two major benefits. First, wall‑clock time is reduced dramatically: the distributed EA achieves an average speed‑up of 6.8× over the single‑machine baseline, with near‑linear scaling (≈85 % efficiency) as the number of nodes increases. Communication and synchronization account for roughly 10 % of total runtime, indicating that the approach remains efficient even for moderately large datasets. Second, the quality of the discovered models improves. Because the cluster can evaluate many more random seeds and hyper‑parameter settings in parallel, the best PUNN found by the distributed search attains on average 2.4 % higher classification accuracy and uses about 17 % fewer parameters than the best model obtained by the single‑machine EA. On the large text corpora the accuracy rises from 92.3 % to 94.7 % while maintaining comparable depth, and the number of hidden units drops by 12 %. Compared with a conventional MLP of equal size, the PUNN delivers a 3.1 % accuracy gain, and although deep CNN/Transformer models still achieve the highest absolute performance, the PUNN offers a superior accuracy‑to‑parameter ratio (20–30 % better).
The authors discuss several limitations. Data transfer, while modest in the current setting, could become a bottleneck for truly massive datasets, suggesting the need for more sophisticated data‑sharding or in‑situ processing. The EA’s performance is sensitive to its own meta‑parameters (mutation rate, crossover probability, population size), and the paper does not explore automated tuning of these settings. Finally, the implementation relies on CPU resources; integrating GPU acceleration for the training phase could further reduce evaluation time and enable exploration of even larger populations.
In conclusion, the study validates that distributing the evolutionary search for PUNNs across a cluster yields both computational efficiency and higher‑quality classification models. The approach is generic and could be applied to other non‑convex model families that require global architecture search, such as symbolic regression or evolutionary circuit design. Future work will extend the framework to hybrid CPU‑GPU clusters, incorporate cloud‑based auto‑scaling, and adopt multi‑objective optimization to simultaneously minimize latency, memory footprint, and energy consumption, thereby making evolutionary design of complex neural architectures practical for real‑time and large‑scale applications.