Factorial graphical lasso for dynamic networks
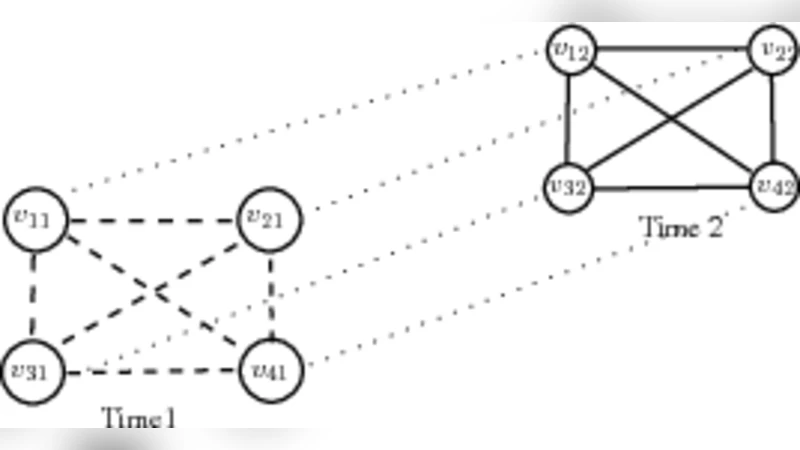
Dynamic networks models describe a growing number of important scientific processes, from cell biology and epidemiology to sociology and finance. There are many aspects of dynamical networks that require statistical considerations. In this paper we focus on determining network structure. Estimating dynamic networks is a difficult task since the number of components involved in the system is very large. As a result, the number of parameters to be estimated is bigger than the number of observations. However, a characteristic of many networks is that they are sparse. For example, the molecular structure of genes make interactions with other components a highly-structured and therefore sparse process. Penalized Gaussian graphical models have been used to estimate sparse networks. However, the literature has focussed on static networks, which lack specific temporal constraints. We propose a structured Gaussian dynamical graphical model, where structures can consist of specific time dynamics, known presence or absence of links and block equality constraints on the parameters. Thus, the number of parameters to be estimated is reduced and accuracy of the estimates, including the identification of the network, can be tuned up. Here, we show that the constrained optimization problem can be solved by taking advantage of an efficient solver, logdetPPA, developed in convex optimization. Moreover, model selection methods for checking the sensitivity of the inferred networks are described. Finally, synthetic and real data illustrate the proposed methodologies.
💡 Research Summary
Dynamic networks—systems whose interactions evolve over time—are central to many scientific domains, from cellular biology and epidemiology to sociology and finance. Estimating the underlying structure of such networks is notoriously difficult because the number of potential edges grows quadratically with the number of nodes, while the number of temporal observations is often limited. Traditional approaches, most notably the Gaussian Graphical Lasso (GGL), impose an ℓ₁ penalty to encourage sparsity but assume a static precision matrix and ignore temporal constraints or prior knowledge about the presence or absence of specific links.
In this paper the authors introduce the Factorial Graphical Lasso (FGL), a structured Gaussian dynamical graphical model that explicitly incorporates three kinds of domain‑driven constraints: (1) known edges (forced to be present or absent), (2) block equality constraints that tie together groups of edges expected to share the same strength across time, and (3) temporal smoothness constraints that limit how much a precision matrix can change from one time point to the next. By encoding these constraints as linear equalities, the number of free parameters is dramatically reduced, turning an otherwise intractable high‑dimensional estimation problem into a convex optimization task.
Mathematically, the objective function is
\
Comments & Academic Discussion
Loading comments...
Leave a Comment