Conformational selection and induced changes along the catalytic cycle of E. coli DHFR
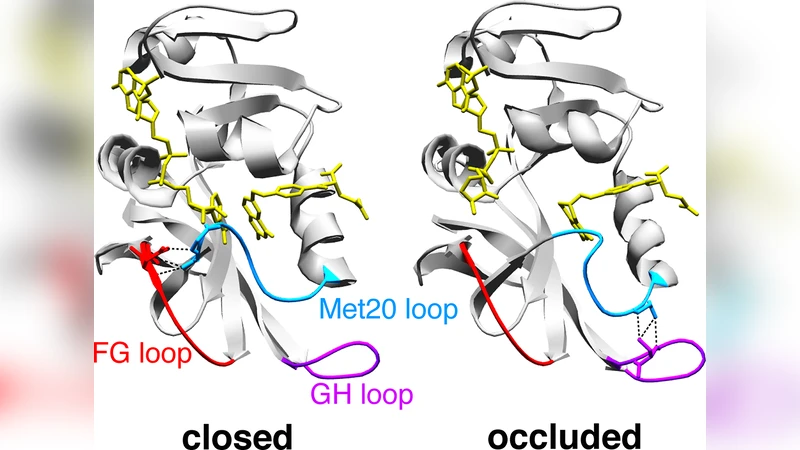
Protein function often involves changes between different conformations. Central questions are how these conformational changes are coupled to the binding or catalytic processes during which they occur, and how they affect the catalytic rates of enzymes. An important model system is the enzyme dihydrofolate reductase (DHFR) from E. coli, which exhibits characteristic conformational changes of the active-site loop during the catalytic step and during unbinding of the product. In this article, we present a general kinetic framework that can be used (1) to identify the ordering of events in the coupling of conformational changes, binding and catalysis and (2) to determine the rates of the substeps of coupled processes from a combined analysis of NMR R2 relaxation dispersion experiments and traditional enzyme kinetics measurements. We apply this framework to E. coli DHFR and find that the conformational change during product unbinding follows a conformational-selection mechanism, i.e. the conformational change occurs predominantly prior to unbinding. The conformational change during the catalytic step, in contrast, is an induced change, i.e. the change occurs after the chemical reaction. We propose that the reason for these conformational changes, which are absent in human and other vertebrate DHFRs, is robustness of the catalytic rate against large pH variations and changes to substrate/product concentrations in E. coli.
💡 Research Summary
The paper presents a comprehensive kinetic framework that integrates NMR relaxation‑dispersion data with classical enzyme‑kinetic measurements to dissect the temporal ordering of conformational changes, ligand binding, and chemical catalysis in Escherichia coli dihydrofolate reductase (DHFR). DHFR catalyzes the reduction of dihydrofolate (DHF) to tetrahydrofolate (THF) using NADPH, a reaction accompanied by a characteristic motion of the Met20 loop that can adopt a “closed” (loop tucked over the active site) or an “open” conformation. The authors first measured the exchange rates between these two loop states using CPMG‑based R₂ relaxation‑dispersion experiments, obtaining forward and reverse rates (k₁ and k₋₁) on the order of 10³–10⁴ s⁻¹. These rates are comparable to, or slightly faster than, the overall catalytic turnover (k_cat ≈ 1.5 × 10³ s⁻¹), indicating that loop motions are fast enough to be coupled to the chemical steps.
Next, they performed traditional kinetic assays (Michaelis–Menten, steady‑state, and pre‑steady‑state measurements) across a range of pH values and substrate/product concentrations to extract the macroscopic rate constants for substrate binding (k_on), product release (k_off), and the chemical step (k_chem). By fitting both data sets simultaneously to a set of mechanistic schemes, they could test two competing models for each stage of the catalytic cycle:
-
Conformational‑selection model for product release – the enzyme first interconverts from the closed to the open loop state (E_closed ⇌ E_open) and only the open form can release THF·NADP⁺. The fitted parameters show that the forward loop opening rate (k₁) exceeds the product‑off rate from the closed form, and that k_off(E_open) ≫ k_off(E_closed). Consequently, product dissociation predominantly occurs after the loop has opened, a classic conformational‑selection scenario.
-
Induced‑fit model for the catalytic step – substrate binding and the chemical reduction (E·NADPH·DHF → E*·NADP⁺·THF) happen while the loop is still in the closed conformation. After the chemical transformation, the loop closes (or re‑closes) in a slower induced transition (k_induced). The kinetic analysis reveals k_chem ≈ 1.5 × 10³ s⁻¹, substantially faster than k_induced (≈ 3 × 10² s⁻¹), indicating that the chemical step precedes the conformational change, i.e., an induced‑fit mechanism.
A comparative discussion with human DHFR highlights that the Met20 loop dynamics observed in E. coli are absent in vertebrate enzymes, which display a more rigid active site and a stronger pH dependence of catalytic rates. The bacterial loop, however, maintains nearly constant exchange rates from pH 5 to pH 9, rendering the overall catalytic rate robust against environmental fluctuations in pH and substrate/product concentrations. The authors argue that this dynamic flexibility provides a selective advantage for E. coli, allowing it to sustain folate metabolism under diverse and rapidly changing growth conditions.
The study’s broader significance lies in its methodological contribution: by coupling site‑specific NMR relaxation data with global kinetic modeling, the authors demonstrate a powerful approach to resolve the sequence of events in complex enzymatic cycles. This framework can be applied to other enzymes where conformational dynamics are suspected to modulate activity, and it offers a rational basis for engineering enzymes or designing inhibitors that target specific conformational states. In summary, the work establishes that DHFR utilizes conformational selection for product release and induced fit for the chemical step, and that these distinct mechanisms together endow the bacterial enzyme with remarkable catalytic robustness.
Comments & Academic Discussion
Loading comments...
Leave a Comment