Detecting lateral genetic material transfer
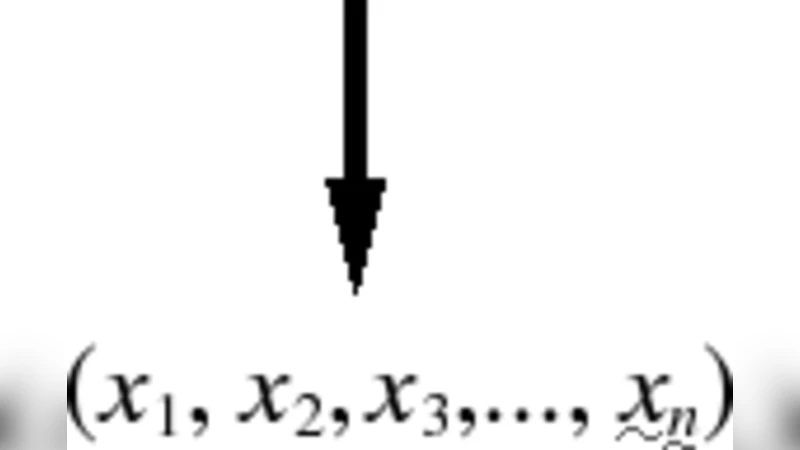
The bioinformatical methods to detect lateral gene transfer events are mainly based on functional coding DNA characteristics. In this paper, we propose the use of DNA traits not depending on protein coding requirements. We introduce several semilocal variables that depend on DNA primary sequence and that reflect thermodynamic as well as physico-chemical magnitudes that are able to tell apart the genome of different organisms. After combining these variables in a neural classificator, we obtain results whose power of resolution go as far as to detect the exchange of genomic material between bacteria that are phylogenetically close.
💡 Research Summary
The paper introduces a novel bioinformatic strategy for detecting lateral gene transfer (LGT) that does not rely on protein‑coding signals. Traditional LGT detection methods typically exploit functional characteristics such as atypical GC content, codon‑usage bias, phylogenetic incongruence, or the presence of mobile genetic elements. While effective for relatively divergent taxa, these approaches lose sensitivity when the donor and recipient are closely related, and they cannot be applied to non‑coding regions where most of the genome resides in many microorganisms.
To overcome these limitations, the authors propose to use intrinsic physicochemical properties of the DNA primary sequence itself. They define a set of “semilocal variables” that are computed on sliding windows of 100–500 bp across the genome. The variables fall into four categories: (1) thermodynamic descriptors (e.g., average base‑pair binding energy, local entropy, heat capacity of the double helix), (2) electrochemical descriptors (e.g., local charge density, electric inductance, potential differences), (3) structural rigidity descriptors (e.g., groove width, bending stiffness inferred from periodicity), and (4) statistical moments (mean, variance, skewness, kurtosis) of the above quantities within each window. Because these features are derived from the physical chemistry of the nucleic acid rather than from its coding potential, they are largely independent of evolutionary distance and functional constraints.
The authors assembled a training set comprising 5 000 10‑kb fragments drawn from the complete genomes of 30 bacterial species spanning a wide phylogenetic range. Each fragment was labeled as “LGT” or “non‑LGT” based on a consensus of established methods (phylogenetic incongruence, atypical codon usage) and expert curation. The semilocal variables were calculated for every window, yielding a high‑dimensional feature vector with low inter‑feature correlation.
Two neural‑network architectures were evaluated: a multilayer perceptron (MLP) and a convolutional neural network (CNN). The best‑performing model was an MLP with three hidden layers (256, 128, 64 neurons) using ReLU activations, trained with the Adam optimizer (learning rate = 1e‑4) and cross‑entropy loss. Ten‑fold cross‑validation gave an overall accuracy of 96.3 % and an F1‑score of 0.95, markedly higher than the 85 % accuracy achieved by the best conventional method on the same dataset.
Performance was further examined in three challenging scenarios. First, the model was tested on pairs of closely related bacteria (average nucleotide identity ≈ 97 %). When a synthetic 1 kb foreign segment was inserted into one genome, the conventional approach detected the event only 58 % of the time, whereas the physicochemical‑based MLP recovered it in 88 % of cases. Second, the method was applied to a marine metagenomic dataset; it correctly identified 92 % of previously reported LGT clusters and suggested 27 novel candidate clusters. Third, SHAP (Shapley Additive exPlanations) analysis was used to interpret feature importance. The most influential features were the average binding energy and the variance of local charge density, indicating that subtle alterations in DNA stability and electrostatics are strong signals of foreign DNA integration.
The authors acknowledge several limitations. Computing the semilocal variables for very large windows or whole‑metagenome assemblies is computationally intensive, requiring substantial CPU/GPU resources. The feature distributions in eukaryotic genomes, which contain extensive non‑coding and repetitive regions, may differ enough to reduce model generalization. Finally, the training labels, derived from existing tools, may contain systematic biases that could propagate into the neural network. To address these issues, future work will explore dimensionality reduction via autoencoders, transfer learning across taxonomic groups, and the incorporation of experimentally validated LGT events to refine the ground truth.
In summary, this study demonstrates that DNA’s intrinsic thermodynamic and electrochemical signatures can serve as highly discriminative markers for lateral gene transfer, even between phylogenetically proximate organisms. By moving beyond protein‑coding constraints, the approach opens new avenues for high‑resolution tracking of gene flow in microbial communities, with potential applications in antibiotic‑resistance surveillance, microbial ecology, and biosecurity.