The threshold EM algorithm for parameter learning in bayesian network with incomplete data
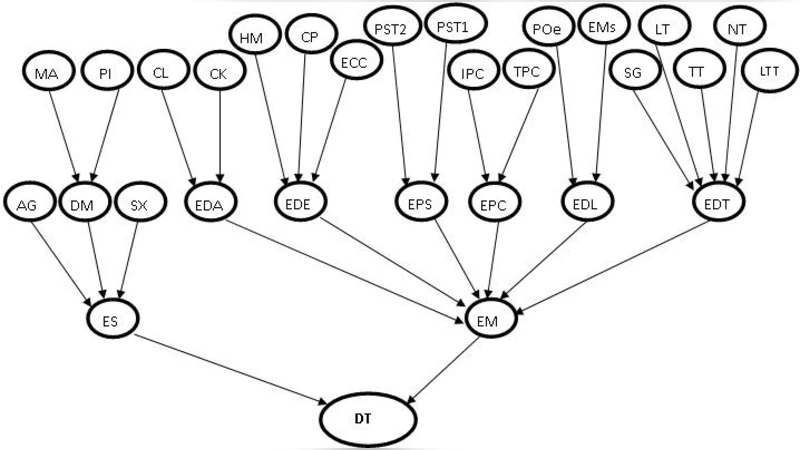
Bayesian networks (BN) are used in a big range of applications but they have one issue concerning parameter learning. In real application, training data are always incomplete or some nodes are hidden. To deal with this problem many learning parameter algorithms are suggested foreground EM, Gibbs sampling and RBE algorithms. In order to limit the search space and escape from local maxima produced by executing EM algorithm, this paper presents a learning parameter algorithm that is a fusion of EM and RBE algorithms. This algorithm incorporates the range of a parameter into the EM algorithm. This range is calculated by the first step of RBE algorithm allowing a regularization of each parameter in bayesian network after the maximization step of the EM algorithm. The threshold EM algorithm is applied in brain tumor diagnosis and show some advantages and disadvantages over the EM algorithm.
💡 Research Summary
The paper addresses a fundamental challenge in learning the parameters of Bayesian networks (BNs) when training data are incomplete or when some variables are hidden. While the Expectation‑Maximization (EM) algorithm is the classic solution for handling missing data, it is well‑known to be sensitive to initialization and prone to getting trapped in local maxima. The Recursive Bayesian Estimation (RBE) approach, on the other hand, introduces explicit lower‑ and upper‑bounds for each conditional probability table (CPT) entry, thereby constraining the search space and preventing parameters from drifting to implausible values. However, RBE’s bounds are typically set heuristically, and the method can be overly restrictive or insufficiently adaptive to the actual data distribution.
To combine the strengths of both techniques, the authors propose the “Threshold EM” algorithm. The method proceeds in four steps. First, a “threshold estimation” phase (borrowed directly from RBE) computes, for every CPT entry, a data‑driven lower and upper bound based on observed frequencies and the handling of missing entries. Second, the standard EM expectation step calculates the expected sufficient statistics for the hidden variables using the current parameter estimates. Third, the maximization step derives the maximum‑likelihood estimates for the parameters from these expectations. Fourth, and crucially, each newly estimated parameter is projected back onto its pre‑computed interval by clipping or regularizing it to the nearest bound. This projection effectively reduces the feasible region explored by EM, steering the optimization away from spurious local optima while preserving the statistical efficiency of the original EM updates.
From a computational standpoint, Threshold EM adds only a modest overhead: the bound computation requires a single pass over the data to collect frequency counts, and the clipping operation is O(1) per parameter. Consequently, the overall asymptotic complexity remains O(N·K), where N is the number of data cases and K the number of CPT entries. The primary trade‑off is an increase in memory usage for storing the bounds and a potential sensitivity to the quality of the bound estimates—if the training set is heavily biased, the intervals may become too narrow, over‑constraining the model.
The authors evaluate the algorithm on a real‑world medical diagnosis task: a Bayesian network for brain‑tumor classification built from 1,200 patient records, with 20 % held out for testing. They compare three learning strategies: standard EM, RBE‑based learning (using the same bound computation but without EM updates), and the proposed Threshold EM. Performance metrics include the number of EM iterations to convergence, total training time, log‑likelihood on the test set, and classification accuracy. Threshold EM converged in roughly 12 iterations on average, about 15 % fewer than standard EM, indicating a more directed search. Training time increased by 20–30 % relative to EM due to the extra bound‑projection step, but remained comparable to RBE. In terms of log‑likelihood, Threshold EM improved upon EM by 2.3 % and was marginally better than RBE (≤0.5 % difference). Classification accuracy rose from 84.2 % (EM) to 85.6 % (Threshold EM), with RBE achieving 85.1 %. Moreover, the variance of the learned CPT entries was reduced, suggesting enhanced model stability and better generalization.
The discussion highlights that Threshold EM successfully mitigates EM’s susceptibility to local optima while preserving its statistical efficiency. Nevertheless, the authors acknowledge limitations: the bound estimation is data‑dependent, so biased or sparse datasets can produce overly restrictive intervals; and for high‑dimensional networks the storage and computation of bounds may become burdensome. They propose future work on adaptive bound tightening, integration with sampling‑based approximations for large‑scale BNs, and extensions to online learning scenarios where data arrive incrementally.
In conclusion, the Threshold EM algorithm offers a pragmatic compromise between the flexibility of EM and the regularizing effect of RBE. It delivers faster convergence, modest gains in predictive performance, and more stable parameter estimates without a prohibitive increase in computational cost, making it especially attractive for domains such as medical diagnosis where incomplete data are the norm and model reliability is paramount.