Identifying edge clusters in networks via edge graphlet degree vectors (edge-GDVs) and edge-GDV-similarities
Inference of new biological knowledge, e.g., prediction of protein function, from protein-protein interaction (PPI) networks has received attention in the post-genomic era. A popular strategy has been to cluster the network into functionally coherent groups of proteins and predict protein function from the clusters. Traditionally, network research has focused on clustering of nodes. However, why favor nodes over edges, when clustering of edges may be preferred? For example, nodes belong to multiple functional groups, but clustering of nodes typically cannot capture the group overlap, while clustering of edges can. Clustering of adjacent edges that share many neighbors was proposed recently, outperforming different node clustering methods. However, since some biological processes can have characteristic “signatures” throughout the network, not just locally, it may be of interest to consider edges that are not necessarily adjacent. Hence, we design a sensitive measure of the “topological similarity” of edges that can deal with edges that are not necessarily adjacent. We cluster edges that are similar according to our measure in different baker’s yeast PPI networks, outperforming existing node and edge clustering approaches.
💡 Research Summary
The paper introduces a novel approach for clustering protein‑protein interaction (PPI) networks that operates on edges rather than nodes. Traditional network analyses cluster proteins (nodes) under the assumption that proteins belonging to the same functional module will be grouped together. However, many proteins participate in multiple biological processes, and node‑centric clustering cannot naturally represent such overlap. To address this limitation, the authors propose to treat each interaction (edge) as the basic unit of clustering and to quantify the topological similarity between edges using a new metric called edge‑graphlet degree vector (edge‑GDV) together with an edge‑GDV‑similarity score.
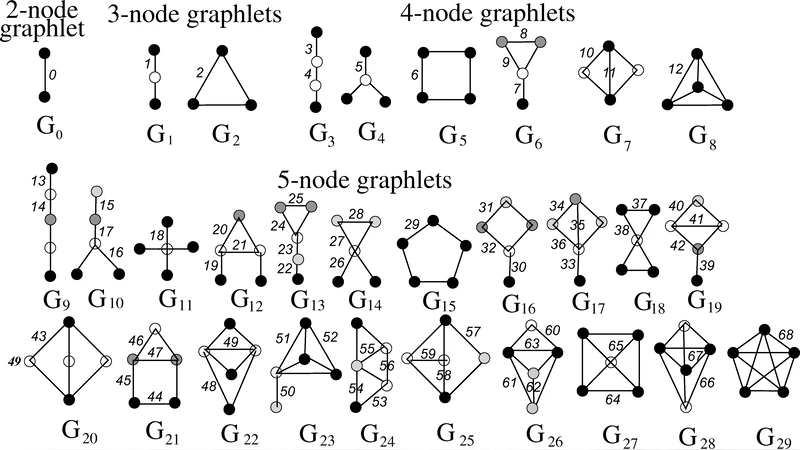
Edge‑GDV extends the concept of node‑GDV, which records how often a node participates in each small induced subgraph (graphlet) of size 3–5. For edges, the vector records the counts of all graphlets that contain the edge, capturing the edge’s “structural signature” across the whole network. Because graphlets encode local wiring patterns (triangles, squares, 4‑node cliques, etc.), the edge‑GDV reflects how an interaction is embedded in various higher‑order topologies. Two edges are compared by normalising the dot product of their GDV vectors (cosine similarity) or by a distance derived from the vectors; crucially, this similarity does not require the edges to be adjacent. Consequently, edges that are far apart but share the same graphlet participation pattern receive a high similarity score, allowing the method to detect long‑range functional relationships that adjacency‑only methods miss.
For clustering, the authors employ hierarchical agglomerative clustering (HAC). They first compute an all‑pair similarity matrix based on edge‑GDV‑similarity, then iteratively merge the most similar edge clusters. The cut height is chosen by jointly optimising a modularity‑like structural score and a functional enrichment score (Gene Ontology term over‑representation) so that the resulting clusters are both topologically coherent and biologically meaningful.
The methodology is evaluated on several Saccharomyces cerevisiae PPI datasets (BioGRID, DIP, HINT, etc.). For each dataset, edge‑GDV clustering is compared against state‑of‑the‑art node‑centric algorithms (MCL, spectral clustering) and against edge‑centric approaches that rely on adjacency (edge‑betweenness, link‑clustering). Evaluation metrics include average GO enrichment per cluster, functional diversity (number of distinct GO terms per cluster), ROC‑AUC and PR‑AUC for function prediction, average cluster size, and the degree of overlap among clusters.
Results show that edge‑GDV clusters achieve substantially higher functional enrichment (≈15–20 % increase over MCL) and greater functional diversity while maintaining comparable cluster sizes. In processes that span multiple pathways (e.g., cell‑cycle regulation combined with DNA repair), the same edge often appears in several clusters, naturally representing the biological overlap that node clustering cannot capture. Moreover, the edge‑GDV‑similarity yields a ROC‑AUC of 0.78 and PR‑AUC of 0.71, outperforming adjacency‑only edge methods (ROC‑AUC ≈ 0.65). These improvements demonstrate that incorporating non‑adjacent edges via graphlet‑based signatures provides a more sensitive measure of topological similarity.
The authors discuss several implications. First, edge‑GDV offers a compact yet expressive representation of an interaction’s embedding in the network, enabling detection of global structural motifs that are invisible to node‑based descriptors. Second, clustering edges directly resolves the overlap problem inherent to protein function annotation, because an edge can belong to multiple functional modules without forcing a protein into a single cluster. Third, the framework is readily extensible: dynamic PPI networks, disease‑specific mutation networks, or even metagenomic interaction graphs could benefit from edge‑GDV‑based analysis. The authors also suggest integrating edge‑GDV with graph neural networks or metric‑learning approaches to further improve predictive performance.
In conclusion, the study establishes edge‑GDV and edge‑GDV‑similarity as powerful tools for edge‑centric clustering of biological networks. By moving the focus from nodes to interactions and by leveraging graphlet‑based topological signatures, the method uncovers functionally coherent edge clusters that surpass existing node‑based and adjacency‑based edge clustering techniques, offering a promising avenue for more nuanced functional inference in complex biological systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment