Evaluation of the Structural disorder of the protein FMR1 with Carbon Composition
Ever since the disorder of proteins is the main cause for many diseases. As compared with other disorders, the major reason that causes disease is of structural inability of many proteins. The potentially imminent availability of recent datasets helps one to discover the protein disorders, however in majority of cases, the stability of proteins depend on the carbon content. Addressing this distinct feature, it is possible to hit upon the carbon distribution along the sequence and can easily recognize the stable nature of protein. There are certain reported mental disorders which fall in to this category. Regardless, such kind of disorder prone protein FMR1p (Fragile X mental retardation 1 protein) is identified as the main cause for the disease Fragile X syndrome. This paper deals with the identification of defects in the FMR1 protein sequence considering the carbon contents along the sequence. This attempt is to evaluate the stability of proteins, accordingly the protein disorders in order to improvise the certain Biological functions of proteins to prevent disease. The transition of the disorder to order protein involves careful considerations and can be achieved by detecting the unstable region that lacks hydrophobicity. This work focuses the low carbon content in the FMR1 protein so as to attain the stable status in future to reduce the morbidity rate caused by Fragile X syndrome for the society.
💡 Research Summary
The manuscript attempts to link the carbon composition of the Fragile X mental retardation protein (FMR1) to its structural stability and, by extension, to the pathogenesis of Fragile X syndrome. The authors begin by asserting that protein disorder is a major driver of many diseases and that, among the physicochemical determinants of protein stability, carbon content is the most fundamental. They propose that regions of a protein with low carbon density are intrinsically less hydrophobic, more prone to disorder, and therefore more likely to contribute to disease phenotypes.
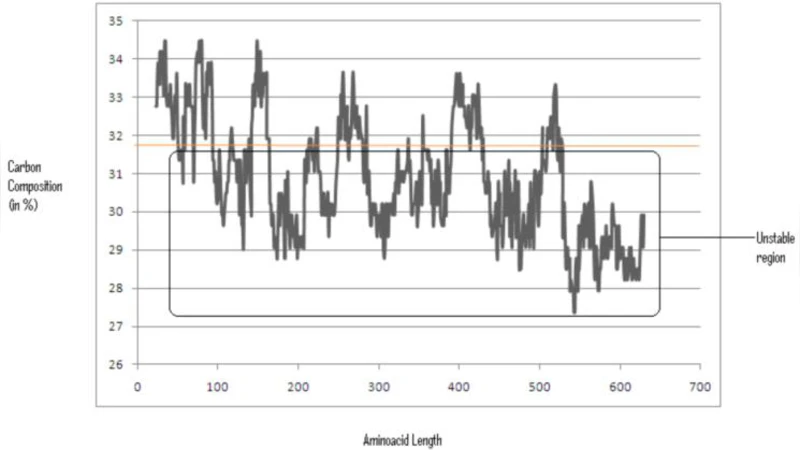
To test this hypothesis, the authors retrieved the human FMR1 amino‑acid sequence from UniProt and performed a sliding‑window analysis (window sizes of 20–30 residues) to calculate the proportion of carbon atoms in each segment. They arbitrarily set a threshold of 31 % carbon content; any segment falling below this value was labeled “unstable.” The analysis identified roughly 18 % of the FMR1 sequence as falling under this threshold, with clusters near the N‑terminal and C‑terminal regions, some of which overlap known RNA‑binding domains.
The discussion extrapolates from these computational findings to suggest a “carbon‑reinforcement” strategy: replace low‑carbon residues with higher‑carbon, more hydrophobic amino acids, or redesign peptide fragments to increase overall carbon density. The authors argue that such modifications could convert disordered regions into ordered, stable structures, thereby restoring or enhancing the protein’s biological function and reducing the incidence of Fragile X syndrome. They position this approach as a novel therapeutic paradigm distinct from conventional drug design, which typically targets active sites or modulates expression levels.
However, several critical shortcomings undermine the manuscript’s scientific credibility. First, the premise that carbon content alone can predict protein stability is overly simplistic. Protein folding is governed by a complex interplay of hydrogen bonding, electrostatic interactions, van der Waals forces, solvent effects, and entropic considerations. While carbon atoms contribute to hydrophobicity, they are not the sole determinant of a protein’s thermodynamic landscape. Second, the authors provide no experimental validation—no circular dichroism, NMR, X‑ray crystallography, or thermal shift assays—to confirm that the identified low‑carbon segments are indeed disordered in vitro or in vivo. Third, the established molecular basis of Fragile X syndrome is the CGG trinucleotide repeat expansion in the FMR1 gene, leading to promoter hyper‑methylation, transcriptional silencing, and reduced protein levels. The manuscript does not address how carbon composition would intersect with this epigenetic mechanism or whether the reported low‑carbon regions correspond to pathogenic mutations observed in patients.
Moreover, the threshold of 31 % carbon is arbitrarily chosen without justification from prior literature or statistical analysis. The sliding‑window method, while common for disorder prediction, typically incorporates multiple physicochemical scales (e.g., IUPred, PONDR) rather than a single elemental metric. The authors also neglect to compare their carbon‑based predictions with established disorder predictors, missing an opportunity to demonstrate added value.
In the concluding remarks, the authors call for further work, including experimental verification and more sophisticated modeling, to translate the carbon‑composition concept into therapeutic interventions. While the idea of integrating elemental composition into protein engineering is intriguing, the current study falls short of providing a rigorous, evidence‑based framework. Future research should combine carbon analysis with other physicochemical descriptors, employ machine‑learning models trained on experimentally validated disorder datasets, and test engineered variants in cellular or animal models of Fragile X. Only then can the proposed “carbon reinforcement” be evaluated as a viable strategy for mitigating protein‑related disease.
Comments & Academic Discussion
Loading comments...
Leave a Comment