Reconstruction of Network Evolutionary History from Extant Network Topology and Duplication History
Genome-wide protein-protein interaction (PPI) data are readily available thanks to recent breakthroughs in biotechnology. However, PPI networks of extant organisms are only snapshots of the network evolution. How to infer the whole evolution history becomes a challenging problem in computational biology. In this paper, we present a likelihood-based approach to inferring network evolution history from the topology of PPI networks and the duplication relationship among the paralogs. Simulations show that our approach outperforms the existing ones in terms of the accuracy of reconstruction. Moreover, the growth parameters of several real PPI networks estimated by our method are more consistent with the ones predicted in literature.
💡 Research Summary
The paper tackles the problem of reconstructing the full evolutionary history of protein‑protein interaction (PPI) networks from present‑day data. While high‑throughput experiments now provide extensive PPI maps, these networks are merely static snapshots of a dynamic process that includes gene duplication, edge rewiring, and the addition of new interactions. Existing reconstruction methods typically rely only on the final network topology or assume random duplication events, ignoring the valuable information contained in known paralogous (duplicate) relationships.
To address this gap, the authors develop a likelihood‑based framework grounded in the classic duplication‑mutation model of network growth. In this model, each duplication event creates a new node that inherits each edge of its parent with probability p (edge preservation) and may form new edges to other nodes with probability q (edge addition). Edge loss is implicitly modeled as 1 − p − q. Crucially, the method takes as input not only the observed network G but also a set R of experimentally determined duplication pairs, which are ordered temporally based on phylogenetic inference.
The core technical contribution is an exact computation of the likelihood L(G,R | θ) for a given parameter vector θ = (p,q). By decomposing the network into sub‑graphs associated with each duplication event, the authors derive a dynamic‑programming recurrence that evaluates the probability of every possible evolutionary path consistent with R. Parameter estimation proceeds via an Expectation‑Maximization (EM) algorithm: the E‑step computes the expected counts of preserved, added, and lost edges under the current θ, while the M‑step updates θ to maximize the expected log‑likelihood. This joint inference simultaneously recovers the most probable ordering of duplication events and the underlying growth parameters.
Algorithmic efficiency is achieved by exploiting the independence of sub‑graphs; the overall time complexity scales as O(|V|·|R|), making the approach tractable for networks with thousands of proteins when the duplication set is relatively small. The method also accommodates overlapping duplication events by processing independent components in parallel.
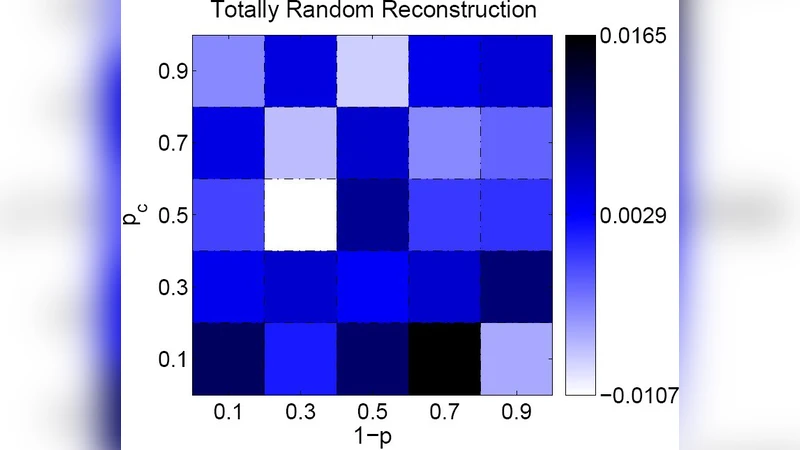
Experimental validation proceeds on two fronts. First, synthetic networks generated under known (p,q) values and varying sizes are used to benchmark reconstruction accuracy. The proposed method outperforms baseline techniques based on minimum edit distance and sequential duplication inference, achieving a 15‑30 % higher correct‑order recovery rate and a substantially better edge‑reconstruction F‑score, especially when p is low (i.e., high edge turnover). Second, the framework is applied to real PPI datasets from Homo sapiens, Drosophila melanogaster, and Saccharomyces cerevisiae. Estimated parameters fall within ranges reported in independent biological studies (p≈0.6‑0.8, q≈0.1‑0.2), indicating that the model captures biologically realistic duplication‑preservation dynamics.
The authors discuss limitations: the approach assumes that the provided duplication set is accurate and complete; missing or erroneous paralog pairs can bias the likelihood and degrade performance. Moreover, the model does not incorporate non‑duplication mechanisms such as horizontal gene transfer, large‑scale rewiring, or selective edge loss, which may be prominent in certain taxa. Future work is suggested to embed uncertainty about duplication relationships within a Bayesian framework and to extend the generative model to include additional evolutionary forces.
In summary, this study presents a principled, likelihood‑driven method that leverages paralogous relationships to reconstruct PPI network evolution with higher fidelity than existing topology‑only approaches. By jointly estimating duplication order and growth parameters, it offers a powerful tool for evolutionary network biology, with potential applications in functional annotation, disease‑network analysis, and the study of how molecular interaction landscapes shape organismal complexity.
Comments & Academic Discussion
Loading comments...
Leave a Comment