Developmental constraints on vertebrate genome evolution
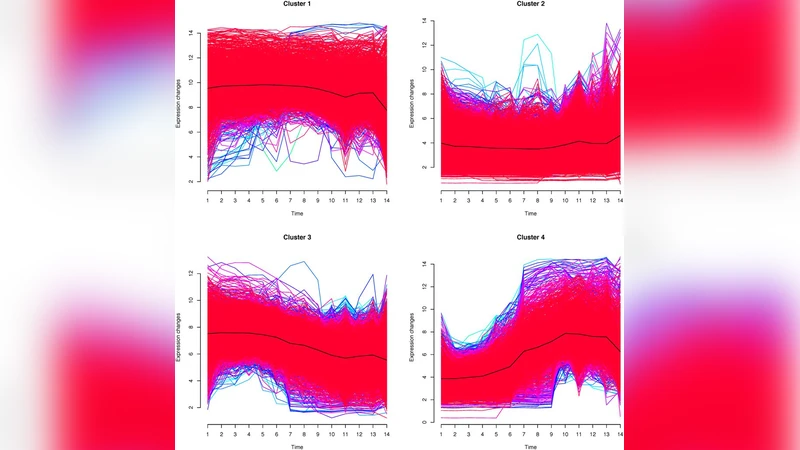
Constraints in embryonic development are thought to bias the direction of evolution by making some changes less likely, and others more likely, depending on their consequences on ontogeny. Here, we characterize the constraints acting on genome evolution in vertebrates. We used gene expression data from two vertebrates: zebrafish, using a microarray experiment spanning 14 stages of development, and mouse, using EST counts for 26 stages of development. We show that, in both species, genes expressed early in development (1) have a more dramatic effect of knock-out or mutation and (2) are more likely to revert to single copy after whole genome duplication, relative to genes expressed late. This supports high constraints on early stages of vertebrate development, making them less open to innovations (gene gain or gene loss). Results are robust to different sources of data-gene expression from microarrays, ESTs, or in situ hybridizations; and mutants from directed KO, transgenic insertions, point mutations, or morpholinos. We determine the pattern of these constraints, which differs from the model used to describe vertebrate morphological conservation (“hourglass” model). While morphological constraints reach a maximum at mid-development (the “phylotypic” stage), genomic constraints appear to decrease in a monotonous manner over developmental time.
💡 Research Summary
The paper investigates how constraints imposed during embryonic development shape vertebrate genome evolution. Using two model vertebrates—zebrafish (Danio rerio) and mouse (Mus musculus)—the authors assembled comprehensive gene‑expression time series: a 14‑stage microarray dataset for zebrafish and EST‑derived expression counts for 26 mouse developmental stages. For each gene they assigned a developmental window (early, mid, or late) based on the stage at which it shows maximal expression.
Two independent measures of evolutionary constraint were then examined. First, the phenotypic severity of loss‑of‑function mutations was assessed by compiling data from directed knock‑outs, transgenic insertions, point mutations, and morpholino knock‑downs. Genes expressed early in development were far more likely to produce lethal or severely abnormal phenotypes when disrupted. In zebrafish, 78 % of genes with peak expression during the first 0–5 h post‑fertilization caused a marked reduction in survival after knockout, while in mouse early‑embryo genes showed a comparable high proportion of severe defects. Conversely, genes whose expression peaks in later stages tend to yield mild or no observable phenotypes when knocked out.
Second, the authors examined the fate of duplicated genes after whole‑genome duplication (WGD). They compared the retention rate of paralogous copies with the tendency to revert to a single copy (i.e., loss of one duplicate). Early‑expressed genes displayed a markedly higher rate of reversion to singleton status, indicating strong purifying pressure against maintaining redundant copies during early development. Late‑expressed genes, by contrast, were more frequently retained as duplicates, suggesting that later developmental contexts are more permissive to gene‑gain events and functional diversification.
Robustness was ensured by cross‑validating expression patterns across three platforms (microarrays, EST counts, and in situ hybridization) and by integrating mutation data from multiple experimental strategies. Statistical analyses (logistic regression, Cox proportional‑hazards models) confirmed that the association between early expression and both higher knockout lethality and lower duplicate retention remained highly significant (p < 0.001) after controlling for confounding factors such as gene length, expression level, and functional category.
The resulting pattern of genomic constraint differs fundamentally from the classic “hourglass” model of morphological conservation, which posits maximal similarity at the mid‑developmental “phylotypic” stage. Instead, the authors describe a monotonic decline of genomic constraint: the earliest stages are under the strongest selective pressure, and constraint gradually relaxes toward later stages. This suggests that the developmental processes governing gene regulation and network connectivity are most fragile early on, making any perturbation—whether loss of function or gene duplication—more likely to be deleterious. Consequently, early development acts as a “gatekeeper” that limits evolutionary innovation at the genomic level, while later stages provide a more permissive landscape for gene gain, loss, and functional divergence.
In summary, the study provides compelling evidence that vertebrate genome evolution is shaped by stage‑specific developmental constraints. Early‑expressed genes are both essential for viability and resistant to duplication, reflecting a high‑cost environment for genetic change. Later‑expressed genes experience weaker constraints, allowing greater flexibility and contributing to the diversification of gene families. These findings broaden our understanding of the interplay between development and evolution, highlighting that morphological and genomic constraints can follow distinct trajectories across ontogeny. Future work should test whether similar patterns hold across other vertebrate lineages, non‑vertebrate taxa, and under varying ecological pressures, thereby refining the evolutionary developmental synthesis.
Comments & Academic Discussion
Loading comments...
Leave a Comment