A Dynamical Model Reveals Gene Co-Localizations in Nucleus
Co-localization of networks of genes in the nucleus is thought to play an important role in determining gene expression patterns. Based upon experimental data, we built a dynamical model to test whether pure diffusion could account for the observed co-localization of genes within a defined subnuclear region. A simple standard Brownian motion model in two and three dimensions shows that preferential co-localization is possible for co-regulated genes without any direct interaction, and suggests the occurrence may be due to a limitation in the number of available transcription factors. Experimental data of chromatin movements demonstrates that fractional rather than standard Brownian motion is more appropriate to model gene mobilizations, and we tested our dynamical model against recent static experimental data, using a sub-diffusion process by which the genes tend to colocalize more easily. Moreover, in order to compare our model with recently obtained experimental data, we studied the association level between genes and factors, and presented data supporting the validation of this dynamic model. As further applications of our model, we applied it to test against more biological observations. We found that increasing transcription factor number, rather than factory number and nucleus size, might be the reason for decreasing gene co-localization. In the scenario of frequency- or amplitude-modulation of transcription factors, our model predicted that frequency-modulation may increase the co-localization between its targeted genes.
💡 Research Summary
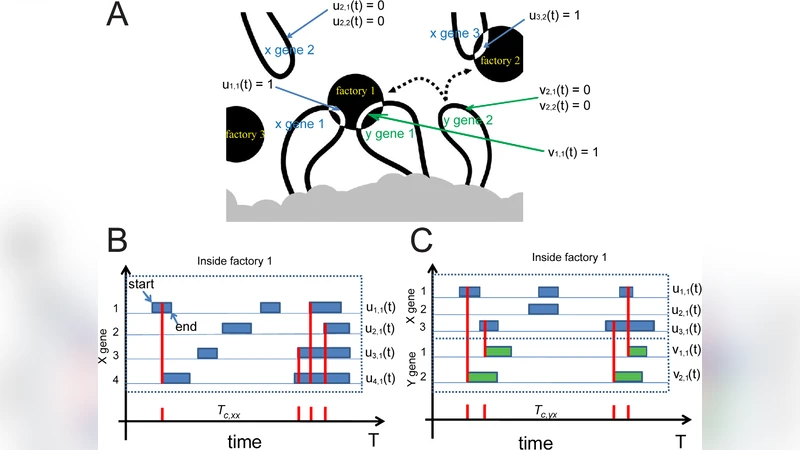
The paper investigates whether simple diffusion can explain the experimentally observed co‑localization of co‑regulated genes within a defined sub‑nuclear region. Starting from live‑cell chromatin‑movement data, the authors first confirm that gene trajectories are better described by fractional (sub‑diffusive) Brownian motion rather than classical Brownian motion. They then construct a stochastic dynamical model in both two‑ and three‑dimensional space in which each gene is represented as a diffusing particle that can transiently bind to a limited number of transcription‑factor “factories” (fixed sites). Binding is treated as a probabilistic collision event; no explicit attractive interaction between genes is required.
Simulation results show that preferential co‑localization emerges naturally when the number of available transcription factors (TFs) is limited. Genes that share the same TF are more likely to be captured by the same factory, leading to spatial clustering. Conversely, increasing the total TF pool reduces the probability of such collisions and weakens co‑localization. Varying the number of factories or the overall nuclear volume has a comparatively minor effect, indicating that TF availability, rather than geometric constraints, is the dominant driver of the phenomenon.
The authors further explore dynamic modulation of TF activity. They distinguish between frequency modulation (rapid on/off switching of TF availability) and amplitude modulation (overall changes in TF concentration). In simulations, frequency‑modulated TFs markedly increase the co‑localization of their target genes, whereas amplitude modulation produces little change. This suggests that temporal patterns of TF signaling can fine‑tune spatial gene organization.
To validate the model, the authors compare its predictions with recent static experimental datasets that measure distances between specific genes and TFs. The predicted co‑localization probabilities correlate strongly with the observed distance distributions, supporting the plausibility of a diffusion‑driven mechanism.
Finally, the model is applied to broader biological questions. It predicts that developmental or environmental conditions that raise TF abundance should lead to a measurable decline in gene co‑localization, while conditions that promote rapid TF cycling (e.g., oscillatory signaling) should enhance clustering. These insights provide a quantitative framework for interpreting how changes in TF number and dynamics can reshape the three‑dimensional genome architecture without invoking direct gene‑gene interactions.
Overall, the study offers a parsimonious yet powerful explanation for gene co‑localization: limited TF resources combined with sub‑diffusive motion are sufficient to generate the observed spatial patterns, and the temporal modulation of TF activity adds an additional layer of regulatory control over nuclear organization.
Comments & Academic Discussion
Loading comments...
Leave a Comment