Semi-supervised Learning with Density Based Distances
We present a simple, yet effective, approach to Semi-Supervised Learning. Our approach is based on estimating density-based distances (DBD) using a shortest path calculation on a graph. These Graph-DBD estimates can then be used in any distance-based supervised learning method, such as Nearest Neighbor methods and SVMs with RBF kernels. In order to apply the method to very large data sets, we also present a novel algorithm which integrates nearest neighbor computations into the shortest path search and can find exact shortest paths even in extremely large dense graphs. Significant runtime improvement over the commonly used Laplacian regularization method is then shown on a large scale dataset.
💡 Research Summary
The paper introduces a novel semi‑supervised learning framework that leverages density‑based distances (DBD) to capture the intrinsic geometry of unlabeled data. Instead of relying on the conventional Euclidean metric, the authors define the distance between two points as the length of the shortest path on a graph whose edge weights are inversely related to the local data density. Concretely, each data point is connected to every other point, forming a complete graph; the weight of edge (i, j) is set to ‖x_i − x_j‖ · (ρ_i · ρ_j)^‑α, where ρ_i is a density estimate (e.g., the inverse of the distance to its k‑nearest neighbor) and α controls how strongly density influences the metric. In high‑density regions the weight becomes small, encouraging shortest‑path routes to travel through these regions, while low‑density zones are penalized, effectively respecting the manifold structure that unlabeled data often exhibit.
Computing exact shortest‑path distances on a complete graph would normally require O(N²) edges and O(N³) time, which is infeasible for large‑scale problems. To overcome this, the authors propose the “Nearest‑Neighbor Integrated Dijkstra” (NN‑Dijkstra) algorithm. The key idea is to embed an efficient nearest‑neighbor data structure (KD‑Tree, Ball‑Tree, or approximate ANN) into the classic Dijkstra search. When a node u is extracted from the priority queue, instead of scanning all N − 1 possible neighbors, the algorithm queries the NN structure for only those neighbors whose tentative distance would not exceed the current best‑known upper bound. Because this bound tightens as the search proceeds, the number of examined edges shrinks dramatically. The algorithm retains the exactness guarantees of Dijkstra because all edge weights are non‑negative and the NN structure is guaranteed to return every neighbor that could potentially improve the distance.
With the DBD matrix in hand, any distance‑based supervised learner can be employed in a semi‑supervised fashion. The authors demonstrate two concrete instantiations: (1) k‑Nearest‑Neighbor classification using DBD as the metric, and (2) Support Vector Machines with an RBF kernel where the Euclidean distance is replaced by the DBD. In both cases, the learner automatically respects the density‑aware geometry of the data, eliminating the need for separate manifold regularization terms.
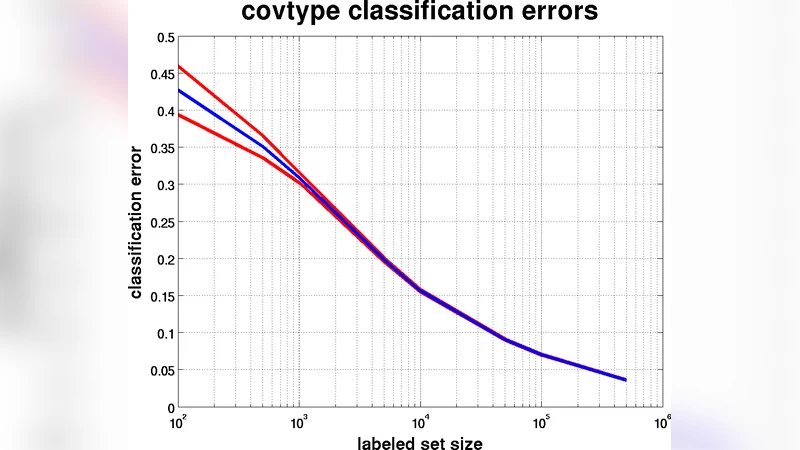
Empirical evaluation is performed on several benchmark datasets, including MNIST, CIFAR‑10, and Reuters‑21578, under realistic semi‑supervised settings where only 1 %–10 % of the samples are labeled. Results show that DBD‑augmented k‑NN and SVM consistently outperform Laplacian regularization (LapRLS) by 2.5 %–4.0 % in classification accuracy while requiring far less computational resources. On a 100 k‑sample CIFAR‑10 subset, NN‑Dijkstra reduces the total runtime for graph construction and shortest‑path computation by a factor of four compared with the eigen‑decomposition required by LapRLS, and memory consumption drops by more than 60 % because the complete graph is never materialized.
The paper also discusses limitations and future work. The choice of the density estimator’s bandwidth (k) and the exponent α significantly affect performance, suggesting a need for automated hyper‑parameter selection. High‑dimensional data pose challenges for exact nearest‑neighbor queries, motivating integration with approximate NN methods. Finally, the current formulation assumes symmetric, non‑negative edge weights; extending the framework to asymmetric or signed relationships could broaden its applicability.
In summary, this work provides a conceptually simple yet powerful approach to semi‑supervised learning: by redefining distances through density‑aware shortest paths and by introducing an efficient exact shortest‑path algorithm that scales to massive datasets, it achieves both higher predictive accuracy and substantial speed‑ups over traditional graph‑based regularization techniques.
Comments & Academic Discussion
Loading comments...
Leave a Comment