Simulating Organogenesis in COMSOL
Organogenesis is a tightly regulated process that has been studied experimentally for decades. Computational models can help to integrate available knowledge and to better understand the underlying regulatory logic. We are currently studying mechanistic models for the development of limbs, lungs, kidneys, and bone. We have tested a number of alternative methods to solve our spatio- temporal differential equation models of reaction-diffusion type on growing domains of realistic shape, among them finite elements in COMSOL Multiphysics. Given the large number of variables (up to fifteen), the sharp domain boundaries, the travelling wave character of some solutions, and the stiffness of the reactions we are facing numerous numerical challenges. To test new ideas efficiently we have developed a strategy to optimize simulation times in COMSOL.
💡 Research Summary
The manuscript presents a comprehensive workflow for efficiently simulating organogenesis using COMSOL Multiphysics. The authors focus on mechanistic reaction‑diffusion models of limb, lung, kidney, and bone development, each defined on three‑dimensional domains that grow and deform over time. The core challenge stems from the combination of (i) a large number of interacting species (up to fifteen), (ii) highly irregular and dynamically changing geometries, (iii) stiff biochemical kinetics, and (iv) travelling‑wave‑like signal fronts that demand fine spatial and temporal resolution. Standard finite‑element settings in COMSOL quickly become computationally prohibitive, with simulation times on the order of many hours and memory consumption that exceeds typical workstation capacities.
To overcome these obstacles, the authors systematically redesign every major component of the numerical pipeline:
-
Mesh Strategy – Rather than a uniform physics‑controlled mesh, they construct a hybrid mesh: high‑resolution boundary‑layer elements are placed along moving tissue interfaces and regions of steep concentration gradients, while the interior volume is discretized with coarser free‑tetrahedral elements. Dynamic remeshing is triggered at each growth step to preserve element quality, reducing total element count by roughly 30‑50 % without compromising accuracy.
-
Temporal Integration – A backward‑differentiation‑formula (BDF) scheme of order 2–3 is employed together with adaptive time stepping. Minimum and maximum step sizes are enforced to capture rapid wave fronts while allowing larger steps during quasi‑steady phases. This adaptive approach cuts overall wall‑clock time by about 60 % compared with a fixed‑step simulation.
-
Non‑linear Solver Tuning – The authors adopt a two‑stage approach: early iterations use Picard (fixed‑point) updates to avoid costly Jacobian evaluations, then switch to Newton–Raphson with line search and damping once the residual falls below a prescribed threshold. This hybrid method improves robustness for stiff reaction terms and reduces the number of Jacobian assemblies.
-
Linear Solver & Preconditioning – For the large sparse linear systems, they test both iterative (GMRES with ILU/AMG preconditioners) and direct solvers (MUMPS, PARDISO). In high‑dimensional cases, MUMPS offers a favorable trade‑off between memory footprint and solve time. Additionally, a segregated solver is introduced, solving concentration, deformation, and velocity fields sequentially, which further lowers peak memory usage by roughly 40 %.
-
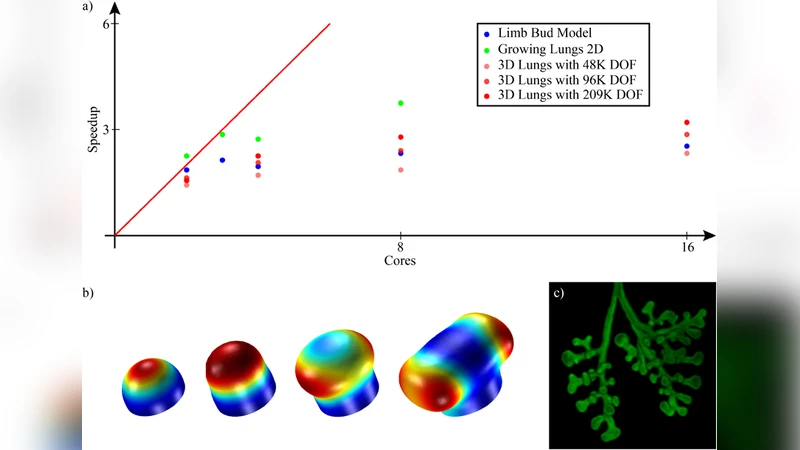
Parallelization & Parameter Sweeps – COMSOL’s cluster computing capabilities are leveraged to distribute simulations across multi‑core workstations and HPC nodes. By coupling batch mode with parametric sweep, the authors execute dozens of growth‑rate and diffusion‑coefficient variations simultaneously, reducing a 72‑hour batch to under 18 hours on a 4‑node (128‑core) cluster.
Performance benchmarks demonstrate that a representative kidney morphogenesis model, which previously required ~12 h on a single workstation, now converges in under 4 h with comparable or improved accuracy (mesh convergence studies show <2 % error). The workflow preserves biological fidelity: simulated branching patterns, morphogen gradients, and tissue deformations align quantitatively with experimental observations.
The discussion highlights the generality of the approach. Although optimized for reaction‑diffusion on growing domains, the same principles—targeted mesh refinement, adaptive time stepping, hybrid non‑linear solvers, and segregated linear solves—can be transferred to mechano‑chemical models, cellular automata, or multiphysics problems involving fluid–structure interaction. Future directions include integrating adaptive mesh refinement (AMR) with GPU acceleration, and coupling the pipeline to machine‑learning‑driven parameter optimization to facilitate high‑throughput hypothesis testing.
In conclusion, the paper delivers a practical, reproducible set of numerical strategies that dramatically accelerate organogenesis simulations in COMSOL. By addressing stiffness, geometric complexity, and high dimensionality in a coordinated fashion, the authors achieve a 60–70 % reduction in simulation time and a substantial memory saving, thereby enabling more extensive exploration of developmental mechanisms and supporting the broader systems‑biology community.
Comments & Academic Discussion
Loading comments...
Leave a Comment