Ensemble Risk Modeling Method for Robust Learning on Scarce Data
In medical risk modeling, typical data are “scarce”: they have relatively small number of training instances (N), censoring, and high dimensionality (M). We show that the problem may be effectively simplified by reducing it to bipartite ranking, and introduce new bipartite ranking algorithm, Smooth Rank, for robust learning on scarce data. The algorithm is based on ensemble learning with unsupervised aggregation of predictors. The advantage of our approach is confirmed in comparison with two “gold standard” risk modeling methods on 10 real life survival analysis datasets, where the new approach has the best results on all but two datasets with the largest ratio N/M. For systematic study of the effects of data scarcity on modeling by all three methods, we conducted two types of computational experiments: on real life data with randomly drawn training sets of different sizes, and on artificial data with increasing number of features. Both experiments demonstrated that Smooth Rank has critical advantage over the popular methods on the scarce data; it does not suffer from overfitting where other methods do.
💡 Research Summary
The paper addresses the challenge of building reliable risk models from “scarce” medical survival data, where the number of observations (N) is small, many observations are censored, and the number of covariates (M) can be comparable to or larger than N. Traditional survival analysis relies on Cox proportional hazards (Cox PH) regression, which models the hazard function over time. Cox PH works well when the number of predictors is modest relative to the number of uncensored events, but it over‑fits on small, high‑dimensional datasets and cannot be applied when M > N. Regularized versions such as the L1‑penalized CoxPath improve variable selection but still struggle when data are extremely scarce.
The authors propose to re‑frame risk prediction as a bipartite ranking problem. They define risk as the probability of experiencing the event before a fixed time threshold T, thereby converting each observation into one of two classes: “early failure” (event before T) or “no early failure” (event after T or censored after T). This binary view reduces the influence of noisy survival times while preserving the ordering information needed for concordance evaluation.
The core contribution is a new ranking algorithm called Smooth Rank. For each feature x_i the method:
- Estimates class‑specific densities g_i1 and g_i2 using a cosine kernel (512 evaluation points).
- Computes a normalized difference
q_i(r) = (g_i1(r) – g_i2(r)) / (π_1·g_i1(r) + π_2·g_i2(r)),
where π_k are class priors. Points where the denominator is very small are discarded to avoid instability. - Smooths q_i(r) with LOESS (locally weighted scatterplot smoothing, degree‑1 polynomial) to obtain a robust univariate predictor e_qi(x).
- Assigns a weight w_i = C‑index(e_qi, outcome) – 0.5. A post‑filtering step shrinks any weight below µ/3 (µ = max_j w_j) to zero, effectively discarding weak predictors without manual threshold tuning.
- Aggregates all predictors into a final score
F(x) = Σ_i w_i·e_qi(x_i) / Σ_i w_i.
This scheme mirrors a weighted Naïve Bayes or weighted voting ensemble, but crucially avoids multivariate optimization, making it highly resistant to the curse of dimensionality. The smoothing step further mitigates sampling noise, and the weight‑shrinkage automatically balances strong and weak features.
Experimental evaluation involved ten real‑world survival datasets (e.g., bone‑marrow transplant, colon cancer, lung cancer, breast cancer, primary biliary cirrhosis, and several lymphoma gene‑expression studies). Sample sizes ranged from 97 to 929, while the number of features varied from 7 to 7399, including cases where M ≫ N. The authors compared Smooth Rank against Cox PH and CoxPath, using the concordance index (C‑index) as the performance metric. They also performed two systematic studies: (a) randomly reducing the training set size to simulate increasing scarcity, and (b) fixing the training size while adding irrelevant features to increase dimensionality.
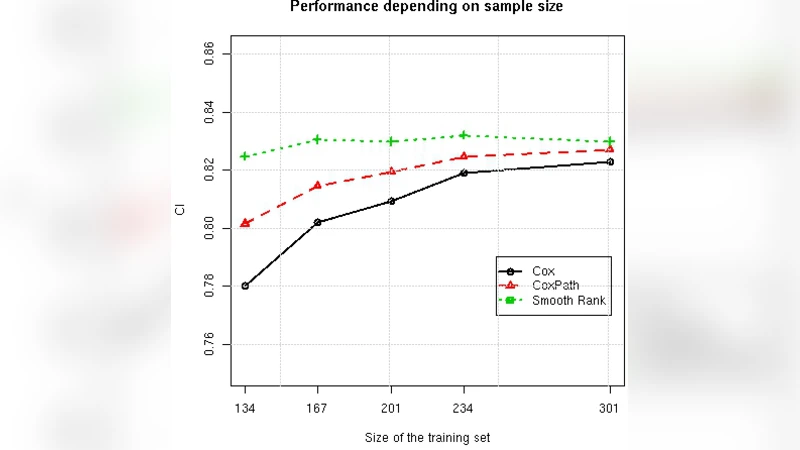
Results: Smooth Rank achieved the highest average C‑index on eight of the ten datasets, losing only on the two datasets with the largest N/M ratios (i.e., the least scarce). In the subsampling experiments, the C‑index of Cox PH and CoxPath dropped sharply as N decreased, whereas Smooth Rank showed a gentle decline, indicating strong resistance to over‑fitting. When the number of features grew, the traditional methods deteriorated rapidly, while Smooth Rank remained comparatively stable. Notably, on gene‑expression datasets where Cox models could not be fitted at all, Smooth Rank still produced meaningful risk scores.
Strengths and limitations: The method’s main advantages are (1) strong regularization via univariate density estimation, (2) robustness from LOESS smoothing, and (3) automatic feature weighting without hyper‑parameter tuning. Potential drawbacks include sensitivity of the density estimates when data are extremely sparse or class imbalance is severe, and the fixed LOESS parameters may not be optimal for every data distribution. Future work could explore adaptive kernel bandwidths and data‑driven smoothing parameters.
Conclusion: By converting survival analysis into a bipartite ranking task and leveraging an ensemble of smoothed univariate predictors, Smooth Rank provides a practical, hyper‑parameter‑free solution that outperforms or matches traditional Cox‑based approaches on scarce, high‑dimensional medical data. This makes it a valuable tool for clinicians and researchers who must build reliable risk models from limited patient cohorts.
Comments & Academic Discussion
Loading comments...
Leave a Comment