Correlating Cell Behavior with Tissue Topology in Embryonic Epithelia
Measurements on embryonic epithelial tissues in a diverse range of organisms have shown that the statistics of cell neighbor numbers are universal in tissues where cell proliferation is the primary cell activity. Highly simplified non-spatial models of proliferation are claimed to accurately reproduce these statistics. Using a systematic critical analysis, we show that non-spatial models are not capable of robustly describing the universal statistics observed in proliferating epithelia, indicating strong spatial correlations between cells. Furthermore we show that spatial simulations using the Subcellular Element Model are able to robustly reproduce the universal histogram. In addition these simulations are able to unify ostensibly divergent experimental data in the literature. We also analyze cell neighbor statistics in early stages of chick embryo development in which cell behaviors other than proliferation are important. We find from experimental observation that cell neighbor statistics in the primitive streak region, where cell motility and ingression are also important, show a much broader distribution. A non-spatial Markov process model provides excellent agreement with this broader histogram indicating that cells in the primitive streak may have significantly weaker spatial correlations. These findings show that cell neighbor statistics provide a potentially useful signature of collective cell behavior.
💡 Research Summary
This paper investigates whether the distribution of cell neighbor numbers (NN) in embryonic epithelia can serve as a quantitative signature of underlying collective cell behavior. The authors begin by confirming that, across a wide range of species (Drosophila, zebrafish, mouse, human, etc.), proliferating epithelia exhibit a remarkably similar NN histogram: most cells have six or seven neighbors, and roughly 90 % of cells possess between four and nine neighbors. This “universal” distribution has previously been attributed to simple, non‑spatial models of cell division.
To test that claim, the authors first implement two classes of non‑spatial models. The first is a discrete‑time Markov chain in which a dividing cell’s two daughters inherit the mother’s neighbor count with a fixed probability. The second is a continuous‑time mean‑field Markov process that treats neighbor number as a globally averaged variable. By sweeping the full parameter space and evaluating the Kullback‑Leibler divergence between model‑generated histograms and the experimental data, they find that both models can only reproduce a narrow slice of the observed distribution and are extremely sensitive to parameter changes. In particular, they cannot simultaneously match the peak at six neighbors and the tails at four and eight neighbors, indicating that ignoring spatial constraints leads to a lack of robustness.
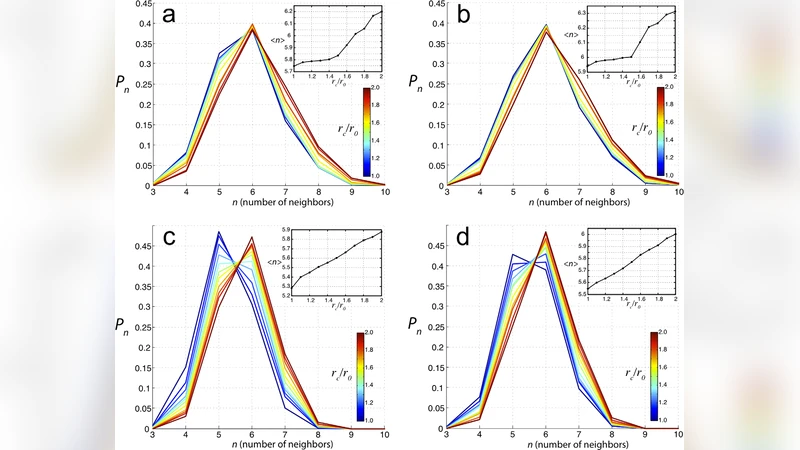
The authors then turn to a spatially explicit approach using the Subcellular Element Model (SCE). In this framework each cell is represented by a collection of interacting particles that enforce volume conservation, cortical tension, and cell‑cell adhesion. Starting from a random polygonal tessellation, cells divide by splitting their area in half, and the mechanical forces naturally rearrange the tissue. Thousands of independent simulations are averaged to produce a steady‑state NN histogram. The resulting distribution aligns closely with the universal experimental histogram, reproducing both the peak position and the relative frequencies of the tails. This success demonstrates that mechanical interactions and local packing constraints are essential for generating the observed statistics.
Having established the importance of spatial correlations for proliferative epithelia, the authors examine a contrasting biological context: the primitive streak region of early chick embryos, where cells actively migrate, intercalate, and ingress. Here the NN histogram is markedly broader, extending from four up to twelve neighbors, and the mean neighbor number is reduced. To capture this behavior, the authors modify the non‑spatial Markov process by allowing daughter cells to be placed at random positions after division, effectively weakening spatial correlations. This adjusted model reproduces the broadened histogram with high fidelity, suggesting that when motility dominates over packing, spatial constraints become less restrictive.
Overall, the study arrives at three key conclusions: (1) non‑spatial proliferation models cannot robustly account for the universal NN distribution observed in proliferating epithelia; (2) spatially resolved mechanical models, exemplified by the SCE, successfully recapitulate the universal histogram, highlighting the role of cell‑cell forces and local geometry; and (3) in tissues where migration and ingression are prominent, a weakened spatial correlation model suffices, and the NN distribution broadens accordingly. The authors propose that the shape of the NN histogram can serve as a diagnostic “signature” of the dominant collective behavior within a tissue.
Finally, the paper outlines future directions, including extending the analysis to three‑dimensional organoids, pathological contexts such as tumor invasion where neighbor statistics may be altered, and the development of image‑analysis pipelines that extract NN histograms from live‑imaging data to infer underlying mechanical or migratory regimes in real time.
Comments & Academic Discussion
Loading comments...
Leave a Comment