Mathematical and computational modeling for describing the basic behavior of free radicals and antioxidants within epithelial cells
The traditional methods of the biology, based on illustrative descriptions and linear logic explanations, are discussed. This work aims to improve this approach by introducing alternative tools to describe and represent complex biological systems. Two models were developed, one mathematical and another computational, both were made in order to study the biological process between free radicals and antioxidants. Each model was used to study the same process but in different scenarios. The mathematical model was used to study the biological process in an epithelial cells culture; this model was validated with the experimental data of Anne Hanneken’s research group from the Department of Molecular and Experimental Medicine, published by the journal Investigative Ophthalmology and Visual Science in July 2006. The computational model was used to study the same process in an individual. The model was made using C++ programming language, supported by the network theory of aging.
💡 Research Summary
The paper addresses a fundamental limitation in traditional biological research on oxidative stress: the reliance on descriptive, linear explanations that cannot capture the inherent non‑linear feedback, saturation, and threshold phenomena of free‑radical (reactive oxygen species, ROS) and antioxidant interactions. To overcome this, the authors develop two complementary quantitative models—a deterministic mathematical model and an agent‑based computational model—and apply each to a different biological context.
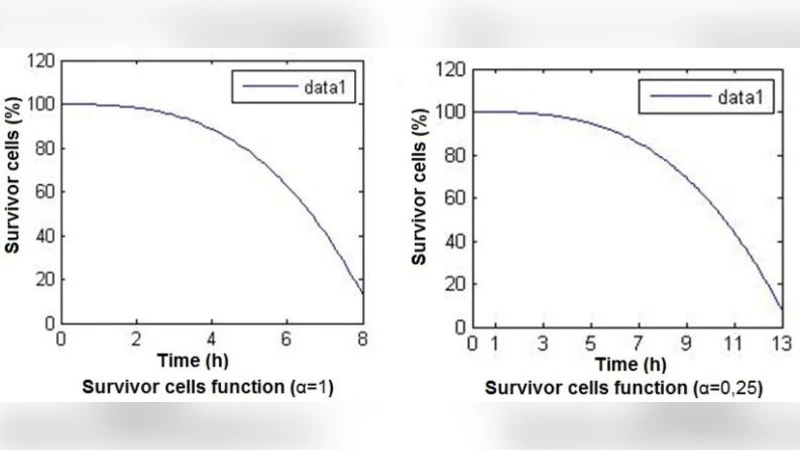
The mathematical model is built from a system of ordinary differential equations (ODEs) describing the time evolution of three state variables: free radicals (R), antioxidants (A), and the radical‑antioxidant complex (C). The equations incorporate radical generation (external stress), radical scavenging by antioxidants, antioxidant regeneration (enzyme‑catalyzed), and natural decay. Non‑linear terms (Michaelis‑Menten kinetics, feedback inhibition) are included to reflect enzyme saturation and the depletion‑induced acceleration of ROS production. Parameter values (reaction rate constants k1–k4, generation rate G, synthesis rate S) are drawn from literature and calibrated against experimental data. Validation uses the 2006 dataset from Anne Hanneken’s group, which measured ROS concentrations over time in cultured ocular epithelial cells. By optimizing parameters to minimize root‑mean‑square error, the model reproduces the observed ROS peak, antioxidant depletion, and subsequent recovery with high fidelity. Sensitivity analysis reveals that modest reductions in key antioxidant enzymes (e.g., superoxide dismutase, catalase) shift the system toward a bifurcation point where ROS explosively increase, highlighting potential therapeutic targets.
The computational model is implemented in C++ as an agent‑based simulation (ABM) that embeds the same ODE dynamics at the cellular level within a spatial network representing tissue architecture. Each cell is an agent (node) possessing its own R and A concentrations, while edges model intercellular signaling and diffusion of oxidative stress. The network topology evolves according to the “network theory of aging”: as chronological age advances, edge weights weaken, and damaged nodes increase the probability of transmitting stress signals to neighbors. Additional parameters capture age‑dependent declines in antioxidant enzyme activity, mitochondrial damage accumulation, and a cell‑death threshold. Simulations of a whole‑organism or organ (e.g., retinal tissue) demonstrate how localized ROS bursts can propagate through weakened connections, leading to systemic oxidative collapse. The model quantifies a critical connectivity density below which the tissue network fragments and ROS levels exceed a global threshold, providing a mechanistic link between cellular aging and organ‑level vulnerability. Importantly, the ABM allows virtual experiments such as targeted antioxidant supplementation in specific regions, revealing that localized boosts can dramatically improve overall network resilience.
By juxtaposing the two approaches, the authors illustrate complementary strengths: the mathematical model excels at precise parameter estimation and direct comparison with bench data, while the computational model captures spatial heterogeneity, network degradation, and emergent tissue‑scale dynamics that are inaccessible to purely analytical methods. Together they form a unified platform for hypothesis testing, enabling in silico trials of antioxidant dosing schedules, enzyme‑enhancing drugs, or lifestyle interventions before costly in vivo studies.
The discussion emphasizes that these quantitative tools move oxidative‑stress research beyond static, linear narratives toward a dynamic systems perspective. The authors propose future extensions, including multi‑scale modeling (cell‑tissue‑organ), integration of omics datasets (transcriptomics, metabolomics) for personalized parameterization, and machine‑learning‑driven optimization of therapeutic protocols. Ultimately, the work demonstrates that mathematically and computationally rigorous models can both validate existing experimental observations and generate novel, testable predictions for combating oxidative damage in epithelial cells and beyond.
Comments & Academic Discussion
Loading comments...
Leave a Comment