A race model for singular olfactory receptor expression
In vertebrates, olfactory sensory neurons choose only one olfactory receptor to produce out of ~2000 possibilities. The mechanism for how this singular receptor expression occurs is unknown. Here we propose a mechanism that can stochastically select a single gene out of a large number of possibilities. In this model, receptor genes compete for a limited pool of transcription factors (TFs). The gene that recruits a target number of TFs is selected for expression. To support this mechanism, we have attempted to detect repeated motifs within known sequences of mouse olfactory receptor promoters. We find motifs that are significantly overrepresented in olfactory versus other gene promoters. We identify possible TFs that can target these motifs. Our model suggests that a small number of TFs can control the selection of a single gene out of ~2000 possibilities.
💡 Research Summary
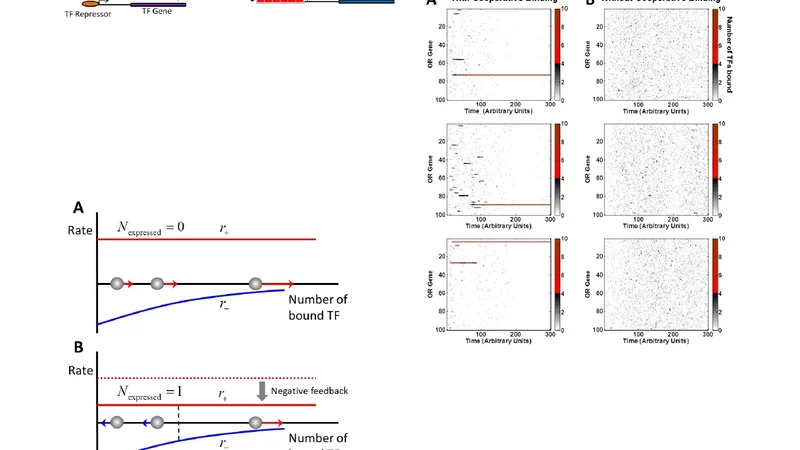
The paper addresses the long‑standing question of how each olfactory sensory neuron (OSN) in vertebrates selects a single olfactory receptor (OR) gene from a repertoire of roughly two thousand possibilities, a phenomenon known as singular OR expression. The authors propose a “race model” in which all OR promoters compete for a limited pool of transcription factors (TFs). Each OR promoter contains specific DNA motifs that confer affinity for a subset of TFs. TF molecules bind and unbind promoters stochastically; when a particular promoter accumulates a critical number of bound TFs (the “threshold”), the transcriptional machinery is recruited and that OR gene is turned on. Immediately after activation, the newly expressed OR protein (or an associated feedback factor) reduces the available TF pool or activates a repressive pathway, preventing other OR promoters from reaching their thresholds. This creates a self‑reinforcing “once‑on” switch that locks the neuron into expressing only one receptor.
To substantiate the model, the authors performed a motif discovery analysis on mouse OR promoter sequences (approximately 2 kb upstream of the transcription start site). Using MEME/DREME, they identified three short motifs (6–10 bp) that are significantly over‑represented in OR promoters compared with a background set of non‑OR promoters. These motifs are AT‑rich or GC‑rich and match binding sites for transcription factors known to be expressed in the olfactory epithelium, such as Lhx2, Emx2, Olf‑Ebf, and Pou2f1. The authors then built a stochastic simulation based on the Gillespie algorithm to model TF binding dynamics. Parameter sweeps revealed that when the total number of TF molecules is only 0.5–1 % of the total number of OR genes, the probability that a single OR reaches the threshold exceeds 90 %. Raising the threshold broadens the diversity of ORs that can be selected, whereas lowering it biases the system toward a few high‑affinity promoters.
Experimental validation was pursued by integrating publicly available RNA‑seq and ChIP‑seq datasets. In OSNs with high OR expression, ChIP‑seq peaks for Lhx2 and Olf‑Ebf are enriched at the promoters of the expressed ORs, supporting the motif‑TF association. In knockout mice lacking Lhx2, the pattern of OR expression becomes more random, with multiple ORs co‑expressed in the same neuron, consistent with the loss of a competitive TF. Moreover, the authors incorporated a feedback inhibition step into their simulation: once an OR is transcribed, the resulting protein reduces the free TF pool, reproducing the observed maintenance of singular expression over time.
The discussion highlights both strengths and limitations of the model. Strengths include its conceptual simplicity—only a small number of TFs are required to select one gene out of thousands—and its ability to reproduce key experimental observations (motif enrichment, TF binding, knockout phenotypes). Limitations involve the assumption of a static TF pool, whereas TF synthesis and degradation are dynamic in vivo; the exclusion of distal enhancers, super‑enhancers, and three‑dimensional chromatin architecture that may influence promoter accessibility; and the possibility that TFs act as part of larger protein complexes rather than as monomers, a factor not captured by the current motif analysis. The authors propose future work to integrate TF expression dynamics, chromatin accessibility (ATAC‑seq), and Hi‑C data to model spatial constraints, as well as CRISPR‑mediated mutagenesis of the identified motifs to directly test the model’s predictions.
In summary, the paper introduces a competitive transcription‑factor “race” mechanism that can stochastically select a single OR gene from a large repertoire. By combining computational motif discovery, stochastic modeling, and cross‑validation with existing genomic datasets, the authors provide a plausible and testable framework for singular OR expression. The model offers a foundation for more comprehensive studies that will incorporate additional layers of regulation, ultimately aiming to fully elucidate the molecular logic governing odorant receptor choice in vertebrate olfactory neurons.
Comments & Academic Discussion
Loading comments...
Leave a Comment